R-TF-028-001 AI Description
Table of contents
- Purpose
- Scope
- Non-Clinical Model Overview
- Description and Specifications
- Integration and Environment
- References
- Traceability to QMS Records
Purpose
This document defines the specifications, performance requirements, and data needs for the non-clinical Artificial Intelligence (AI) models used in the Legit.Health Plus device.
Scope
This document details the design and performance specifications for all non-clinical AI algorithms integrated into the Legit.Health Plus device. It establishes the foundation for the development, validation, and risk management of these models.
This description covers the following key areas for each algorithm:
- Algorithm description, clinical objectives, and justification.
- Performance endpoints and acceptance criteria.
- Specifications for the data required for development and evaluation.
- Requirements related to cybersecurity, transparency, and integration.
- Links between the AI specifications and the overall risk management process.
Non-Clinical Model Overview
The Legit.Health Plus device integrates several non-clinical AI models that are essential for robust, equitable, and high-quality operation of the system. These models do not provide clinical diagnostic outputs, but instead perform technical, quality, and contextual functions that support the overall performance, safety, and fairness of the device. Non-clinical models include:
- Image quality and preprocessing models (e.g., color correction)
- Contextual attribute models (e.g., skin tone identification, body site identification)
- Technical validation models (e.g., 3D reconstruction for area quantification)
These models:
- Perform quality assurance, preprocessing, and technical validation
- Enable downstream clinical models to operate within validated domains and with standardized inputs
- Support equity, bias mitigation, and performance monitoring across diverse populations
- Provide structured, non-clinical metadata (e.g., skin tone, body site, image quality) to enhance device reliability and fairness
- Do not generate clinical diagnostic or severity outputs, nor do they provide interpretative distributions of ICD categories
Key Non-Clinical Models and Their Functions:
- Acneiform Inflammatory Pattern Identification: Translates objective lesion counts and density into standardized IGA severity scores, supporting consistent acne severity assessment.
- Skin Tone Identification: Automatically classifies images by Fitzpatrick and Monk skin tone scales to support bias mitigation, personalization, and regulatory compliance.
- Body Site Identification: Detects anatomical regions present in images, enabling context-aware processing, BSA calculations, and site-specific workflow optimization.
- 3D Surface Area Quantification: Transforms 2D image segmentations into real-world 3D measurements, supporting accurate, reproducible area and volume calculations for research and quality assurance.
- Color Correction: Standardizes color representation in images using reference markers, ensuring reliable color features for downstream models and human interpretation.
These non-clinical models are described in detail in the following section.
Description and Specifications
Acneiform Inflammatory Pattern Identification
Description
A mathematical equation ingests the tabular features derived from the Acneiform Inflammatory Lesion Quantification algorithm and outputs an score, on the scale [4], aligned with Investigator's Global Assessment (IGA).
The equation, with parameters and , takes as input:
- The number of acneiform inflammatory lesions .
- The density of acneiform inflammatory lesions .
The final output is weighted x2.5 to align with an [10] scale, rather than [4], for a more granular output.
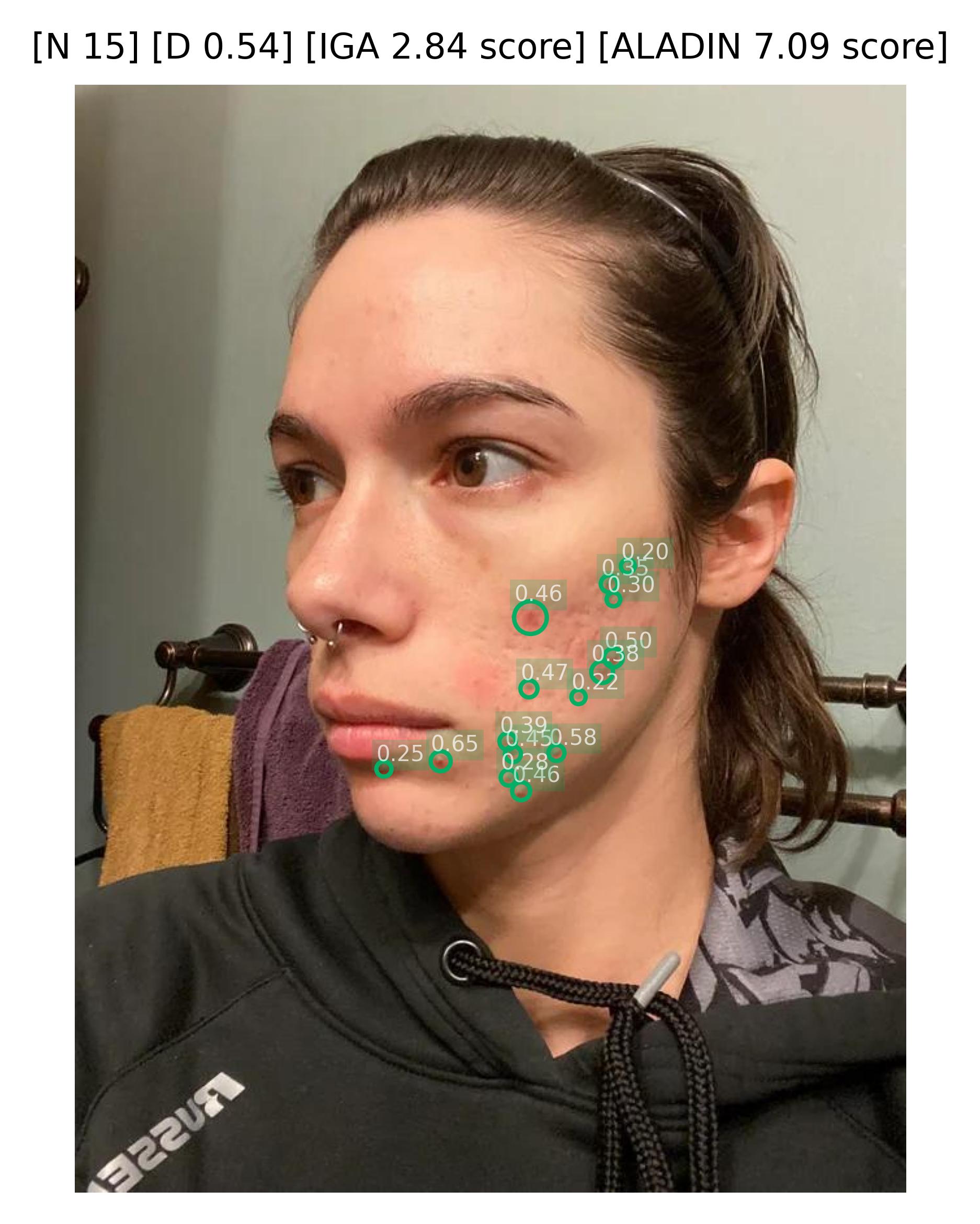 | 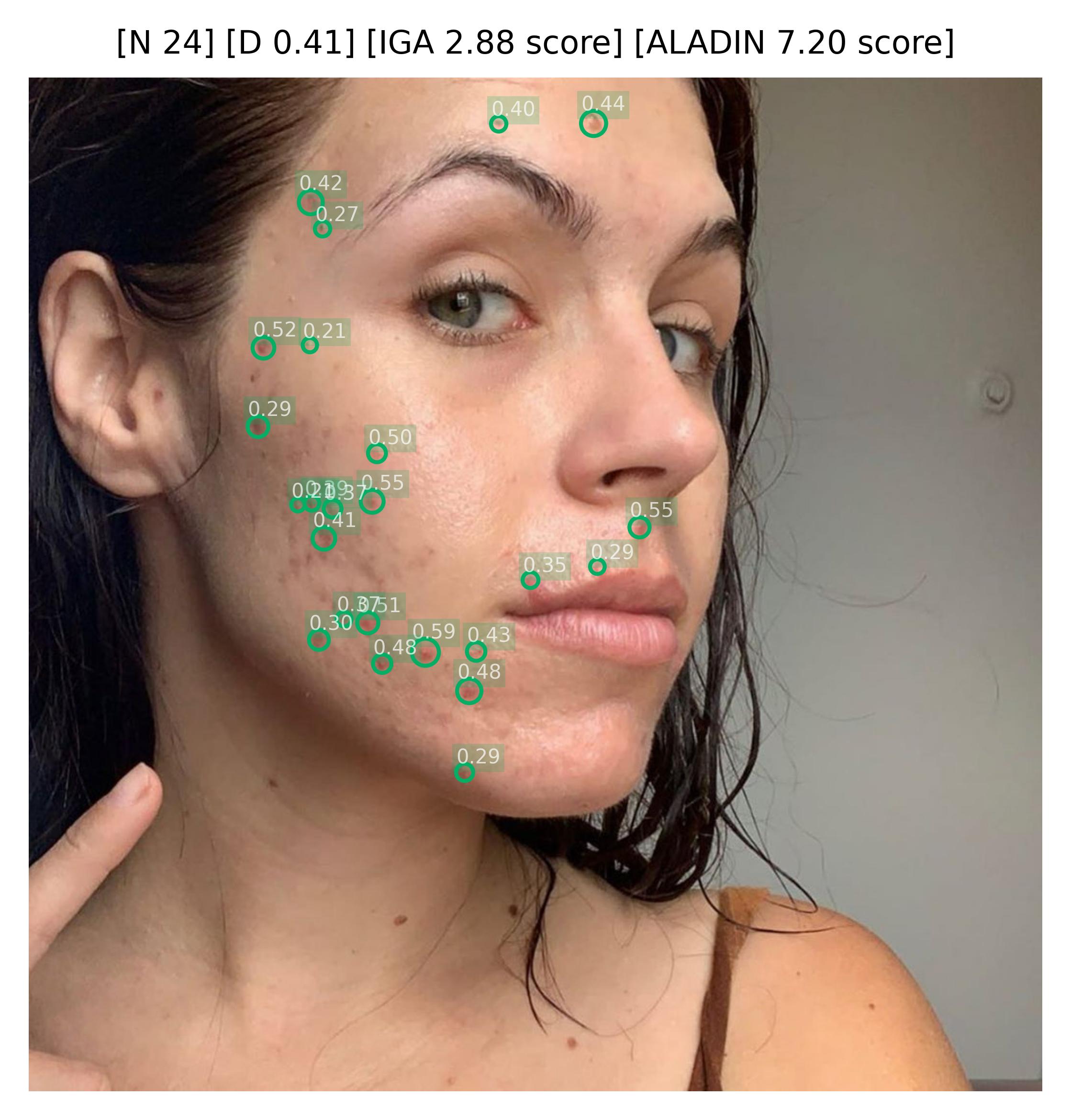 | 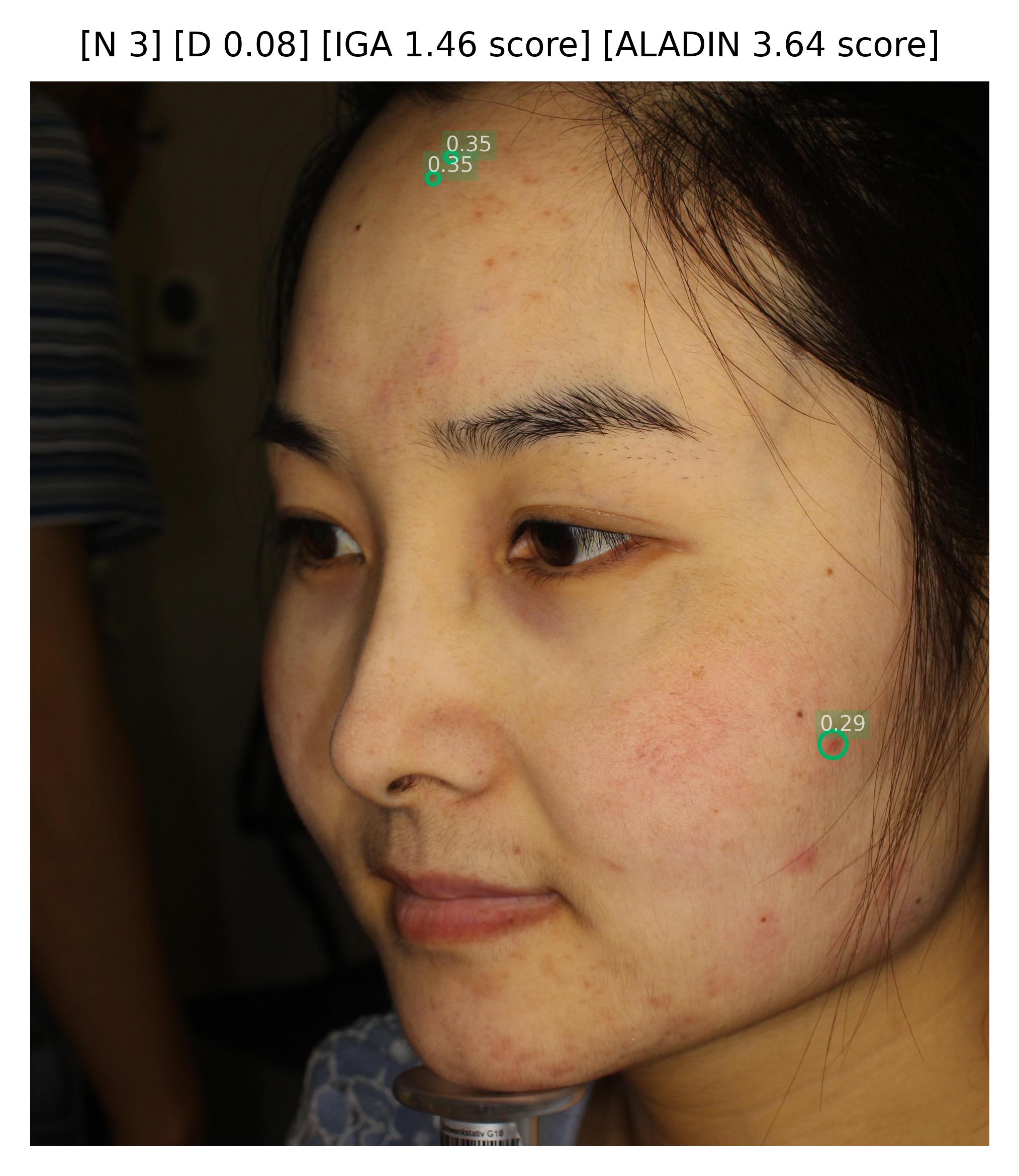 |
|---|
Sample images with acneiform inflammatory lesion detections and their confidence,the number of lesion (N), the density of the lesions (D), the calculated IGA scores, and the calculated ALADIN scores.
Objectives
- Support healthcare professionals in providing standardized acne severity assessment using the validated Investigator's Global Assessment (IGA) scale.
- Reduce inter-observer variability in IGA scoring, which shows moderate agreement (κ = 0.50-0.70) between raters in clinical practice [112].
- Enable automated severity classification by translating objective lesion counts and density into clinically meaningful IGA categories.
- Ensure reproducibility by basing severity assessment on quantitative features rather than subjective visual impression.
- Facilitate treatment decision-making by providing standardized severity grades that align with evidence-based treatment guidelines (e.g., topical therapy for mild, systemic therapy for severe).
- Support clinical trial endpoints by providing consistent, reproducible IGA assessments as required by regulatory agencies.
Justification (Clinical Evidence):
- The IGA scale is a widely validated tool for acne severity assessment and is the most commonly used primary endpoint in acne clinical trials [113, 114].
- Manual IGA assessment shows substantial inter-observer variability (κ = 0.50-0.70), with particular difficulty in distinguishing between adjacent grades [112].
- Objective lesion counting combined with algorithmic severity classification has been shown to improve consistency (κ improvement to 0.75-0.85) compared to purely visual IGA assessment [115].
- Treatment guidelines are explicitly linked to IGA grades, with clear recommendations for topical monotherapy (IGA 1-2), combination therapy (IGA 2-3), and systemic therapy consideration (IGA 3-4) [116].
- Regulatory agencies require validated severity measures for acne trials, with IGA being the most accepted scale for primary efficacy endpoints [117].
- Studies show that automated severity grading reduces assessment time by 40-60% while maintaining or improving accuracy compared to manual grading [118].
Endpoints and Requirements
Performance is evaluated using pearson correlation metric between the predicted and expert consensus, to ensure that the model aligns with the criteria from expert dermatologists.
| Metric | Threshold | Interpretation |
|---|---|---|
| Pearson correlation | ≤ Expert Inter-observer Variability | Model performance is non-inferior to expert inter-observer variability |
Justification of the succeed criteria:
- IGA is the scoring system recommended by the FDA for acne severity assessment in clinical trials. Therefore we seek for high correlation to this scale.
- IGA is inherently subjective, with documented inter-observer variability among dermatologists.
- The stablished succeed criteria ensures that the model's predictions are not less reliable than those made by expert dermatologists, making it suitable for clinical and research applications.
Requirements:
- Implement a tabular model (e.g., gradient boosting, mathematical equation, random forest, neural network, or other ML model) that:
- Accepts numerical inputs dereived from Acneiform Inflammatory Lesion Quantification, such as the total inflammatory lesion count, lesion density, anatomical site identifiers, affected surface area, etc.
- Outputs a severity score highly correlated to the IGA scale.
- Demonstrate a correlation with the ground-truth data non-inferior to the inter-observer variability among expert dermatologists.
- Report all metrics with 95% confidence intervals.
- Validate the model on an independent and diverse dataset including:
- Full range of IGA grades (0-4)
- Diverse patient populations (e.g., various Fitzpatrick skin types)
- Ensure outputs are compatible with:
- FHIR-based structured reporting for interoperability
- Clinical decision support systems for acne treatment recommendations
- Treatment guidelines that specify interventions based on IGA grade
- Clinical trial data collection systems requiring standardized IGA assessments
- Document the model optimization strategy including:
- Feature design
- Hyperparameter optimization methodology
- Rationale for model selection (if multiple architectures compared)
- Provide evidence that:
- The model generalizes across different patient populations
- Predictions align with dermatologist consensus and clinical treatment guidelines
Skin Tone Identification
Description
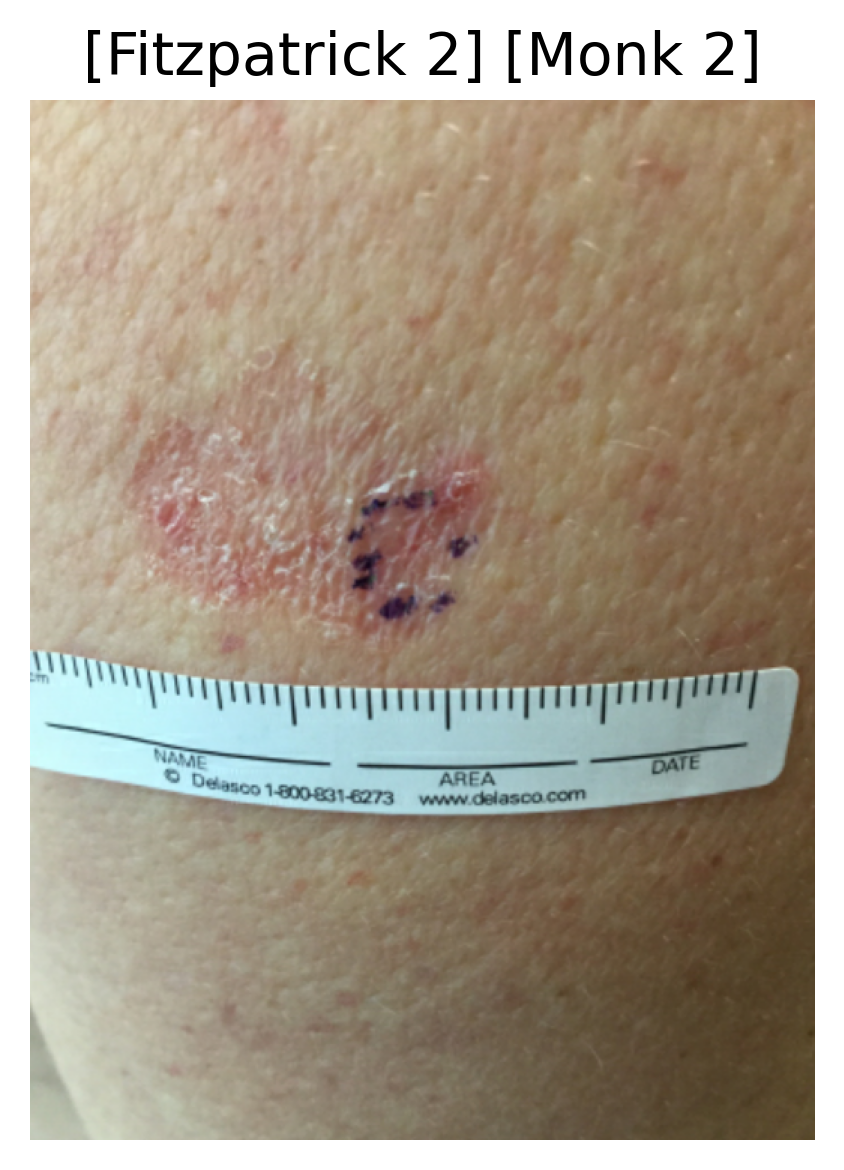
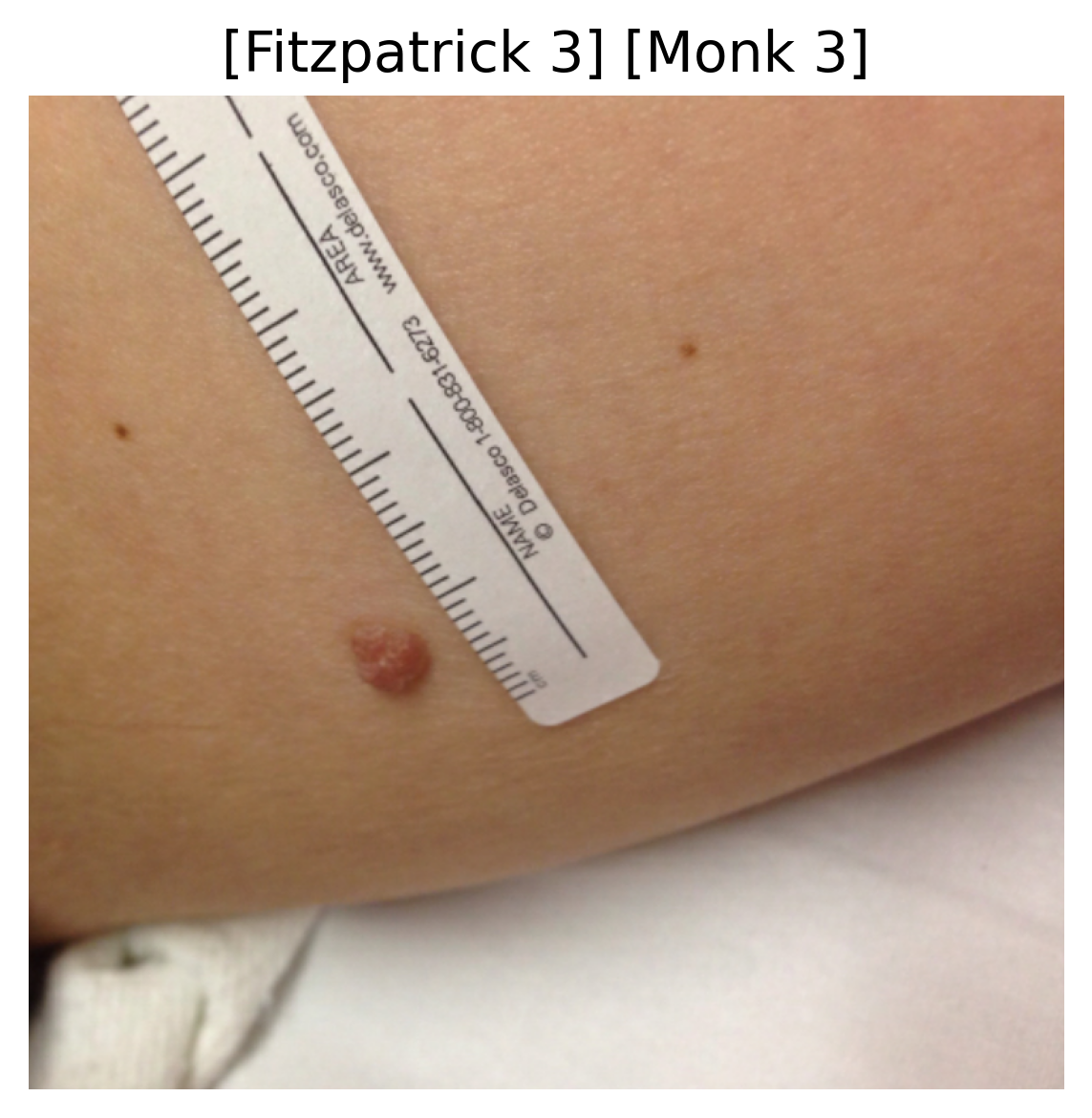
A deep learning multi-class classification model ingests a clinical dermatological image and outputs two probability distributions: one for the six Fitzpatrick skin tone categories and another for the ten Monk skin tone categories.
Fitzpatrick Skin Tone Classification
where each corresponds to the probability that the skin in the image belongs to Fitzpatrick skin tone , and .
The Fitzpatrick skin tones are defined as:
- Type I: Very fair skin, always burns, never tans (pale white skin, often with red/blonde hair)
- Type II: Fair skin, usually burns, tans minimally (white skin, burns easily)
- Type III: Medium skin, sometimes burns, tans uniformly (cream white skin, burns moderately)
- Type IV: Olive skin, rarely burns, tans easily (moderate brown skin)
- Type V: Brown skin, very rarely burns, tans very easily (dark brown skin)
- Type VI: Dark brown to black skin, never burns, tans very easily (deeply pigmented dark brown to black skin)
The predicted Fitzpatrick type is:
Monk Skin Tone Classification
where each corresponds to the probability that the skin in the image belongs to Monk skin tone category , and .
The Monk skin tone categories are defined as:
- Category 0: Lightest skin tone
- Category 1: Very light skin tone
- Category 2: Light skin tone
- Category 3: Light-medium skin tone
- Category 4: Medium skin tone
- Category 5: Medium-dark skin tone
- Category 6: Dark skin tone
- Category 7: Very dark skin tone
- Category 8: Deeply dark skin tone
- Category 9: Darkest skin tone
The predicted Monk skin tone is:
Additional Outputs
For both Fitzpatrick and Monk classifications, the model outputs a confidence scores representing the certainty of the classification.
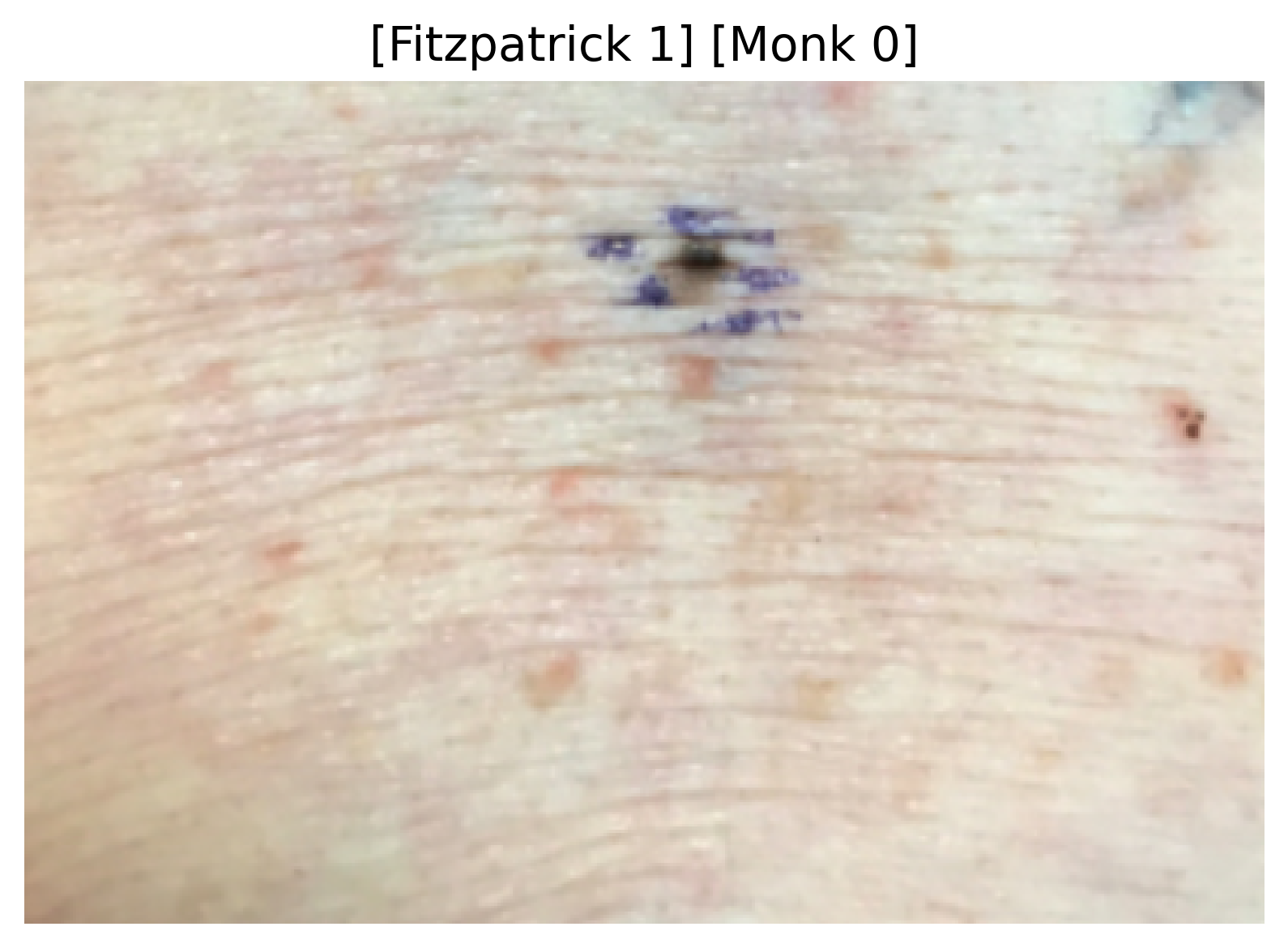 |  |  |
|---|---|---|
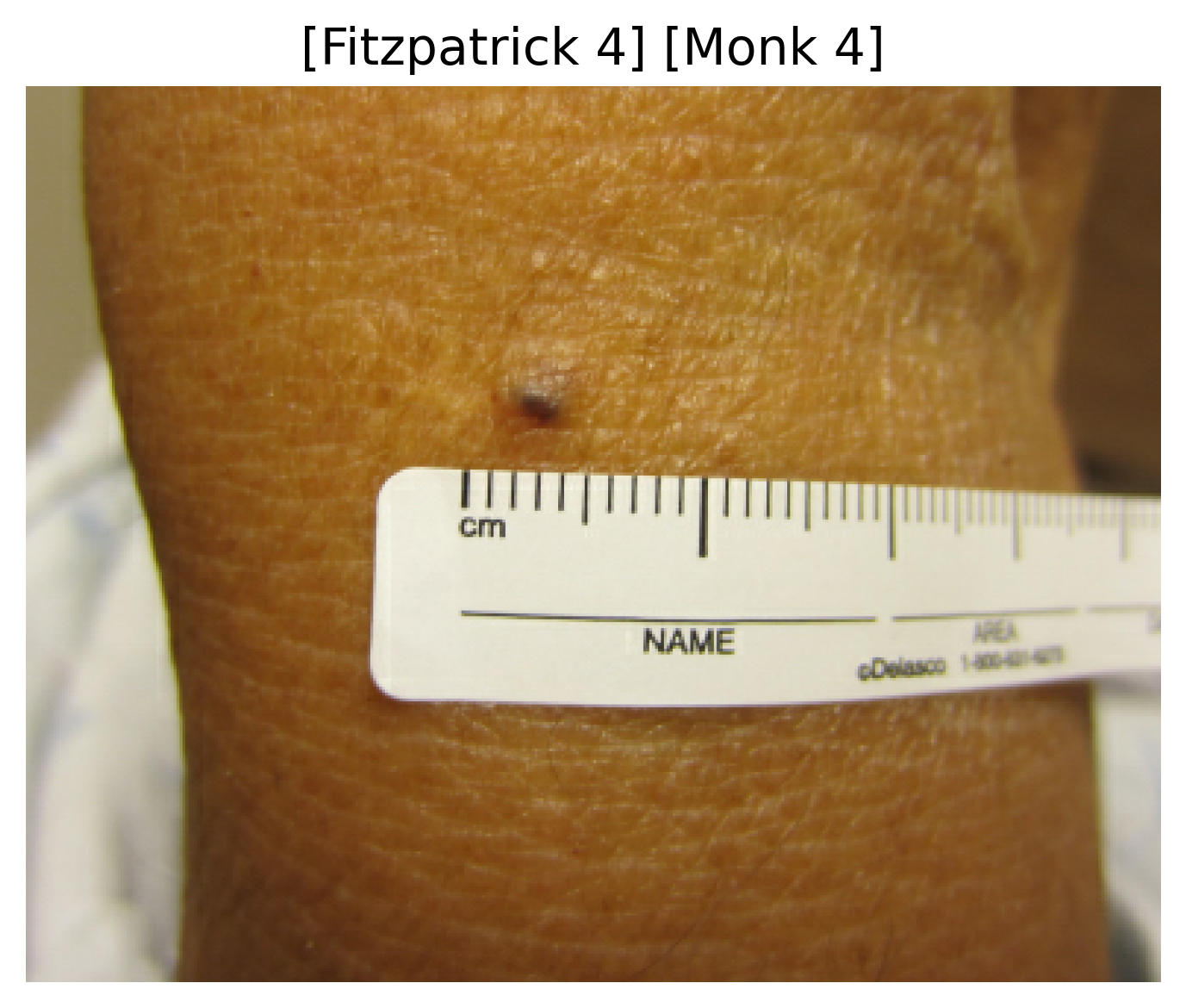 | 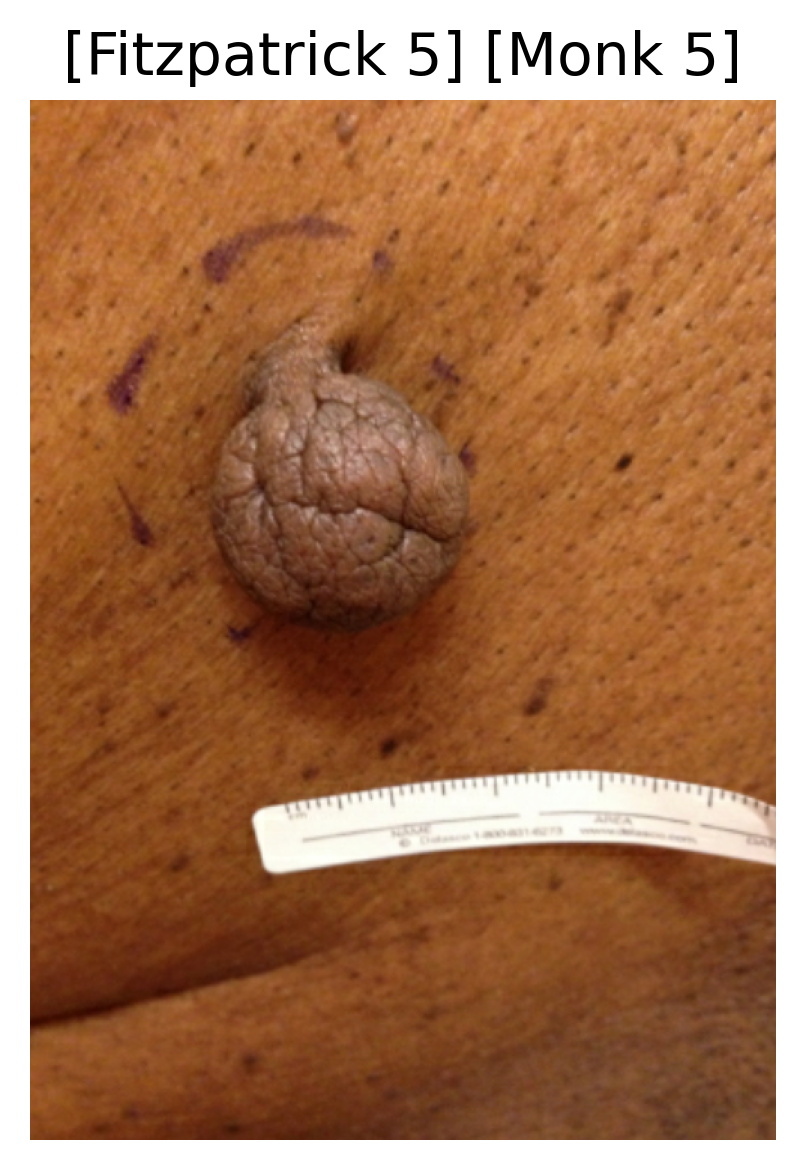 | 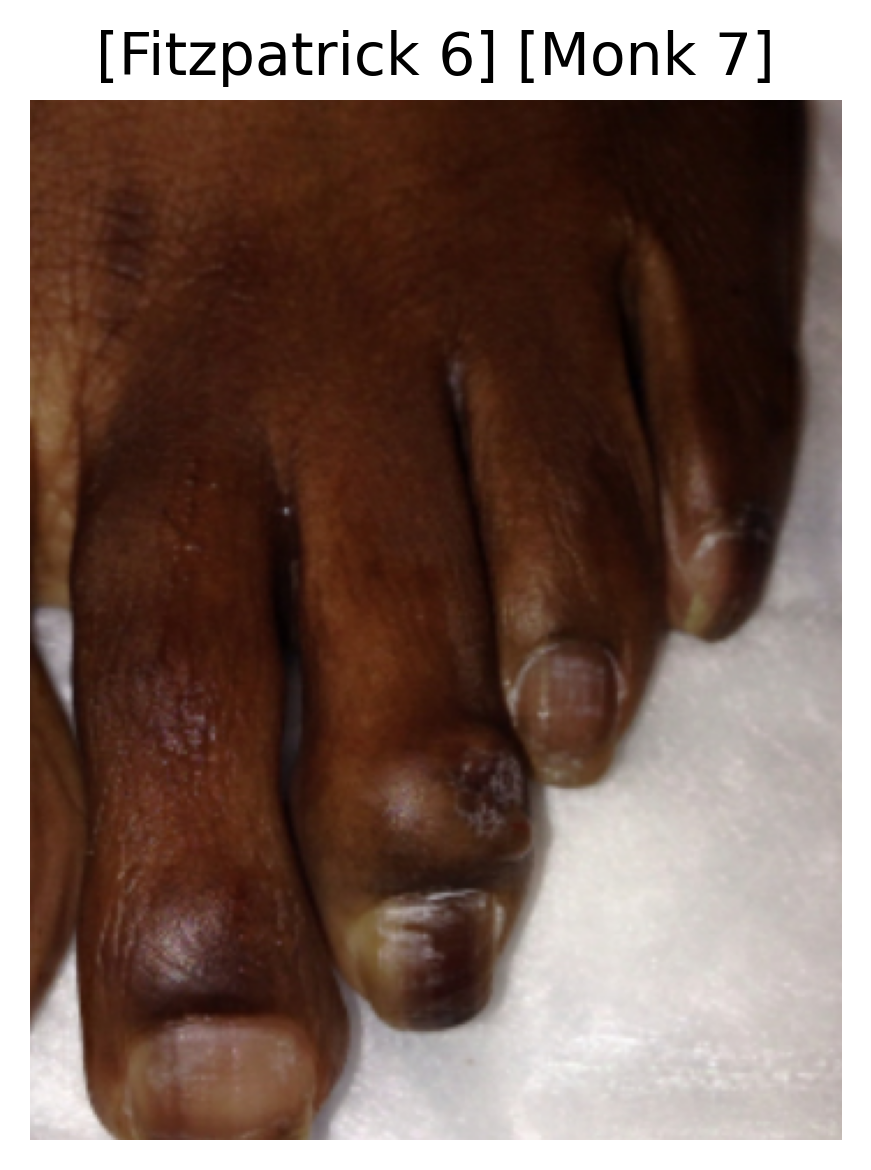 |
Sample images with their predicted Fitzpatrick and Monk skin tone categories.
Objectives
- Enable automated skin tone detection to support personalized dermatological AI models that require skin tone information for accurate predictions.
- Reduce assessment variability in skin tone classification, which shows moderate inter-observer agreement (κ = 0.50-0.65) even among dermatologists [213, 214].
- Support bias mitigation in AI models by identifying underrepresented skin tones in datasets and ensuring equitable performance across all Fitzpatrick and Monk skin tones.
- Facilitate treatment personalization by providing objective skin tone information relevant for phototherapy dosing, laser treatment parameters, and topical therapy selection.
- Enable research stratification by providing consistent skin tone classification for clinical trials and real-world evidence studies.
- Support regulatory compliance by ensuring AI models are validated across diverse skin tones as required by regulatory guidelines.
- Improve telemedicine accessibility by providing automated skin tone assessment in remote settings where patient-reported skin tone may be unreliable.
Justification (Clinical Evidence):
- Fitzpatrick skin tone is a critical factor in dermatological assessment, influencing disease presentation, treatment selection, and AI model performance [215, 216].
- Self-reported Fitzpatrick type shows poor accuracy, with concordance to expert assessment ranging from 40-60%, particularly for intermediate types (III-IV) [217, 218].
- AI model performance shows significant disparities across skin tones, with accuracy degradation of 10-30% for darker skin tones (V-VI) when models are trained on predominantly lighter skin datasets [219, 220].
- Automated skin tone detection enables adaptive AI models that adjust prediction thresholds or use skin tone-specific models, improving accuracy by 15-25% for underrepresented groups [221].
- Treatment dosing for phototherapy and laser procedures requires accurate skin tone assessment, with misclassification leading to suboptimal efficacy or adverse events in 15-20% of cases [222].
- Clinical trials increasingly require Fitzpatrick type stratification to demonstrate equitable treatment efficacy and safety across diverse populations [223].
- Studies show that objective skin tone classification improves inter-rater reliability from κ = 0.50-0.65 (manual) to κ = 0.75-0.85 (automated) [224].
- Automated detection addresses the limitation of visual assessment under different lighting conditions, which can shift perceived skin tone by 1-2 categories [225].
Endpoints and Requirements
Performance is evaluated using classification accuracy and mean absolute error compared to expert Fitzpatrick and Monk assessments.
| Metric | Threshold | Interpretation |
|---|---|---|
| Fitzpatrick Accuracy | ≥ inter-rater variability | Performance non-inferior to expert criteria. |
| Fitzpatrick MAE | ≤ 1 | Average error less than a category away from expert criteria. |
| Monk Accuracy | ≥ inter-rater variability | Performance non-inferior to expert criteria. |
| Monk MAE | ≤ 1 | Average error less than a category away from expert criteria. |
All thresholds must be achieved with 95% confidence intervals.
Threshold Justification:
- The difficulty of skin tone assessment varies significantly between clinical and non-clinical settings.
- The difficulty of skin tone assessment depends on the illuminance quality of the images, which varies significantly between datasets.
- Therefore, it is more appropriate to evaluate model performance against the inter-rater variability established for the specific evaluation dataset, rather than a fixed absolute threshold.
- The ordinal nature of skin tone categories means that adjacent-type misclassifications are more clinically acceptable than distant errors. Therefore, a Mean Absolute Error (MAE) of ≤ 1 category allows for acceptable errors in the vicinity of the true stage, reflecting real-world clinical variability.
Requirements:
- Implement a deep learning classification architecture optimized for skin tone analysis.
- Output structured data including:
- Probability distribution across all categories
- Predicted skin tone with confidence score
- Demonstrate performance meeting or exceeding all thresholds:
- Overall accuracy ≥ inter-rater variability
- MAE ≤ 1
- Ensure outputs are compatible with:
- Downstream AI models that require skin tone information as input
- FHIR-based structured reporting for interoperability
- Clinical decision support systems for treatment personalization
- Bias monitoring dashboards tracking AI performance across skin tones
- Research data collection systems for clinical trial stratification
- Document the training strategy including:
- Data collection protocol ensuring balanced representation
- Multi-expert annotation protocol for ground truth establishment
- Handling of class imbalance (if present)
- Data augmentation strategies preserving skin tone characteristics
- Regularization and calibration techniques
- Transfer learning approach (if applicable)
- Provide evidence that:
- The model provides equitable performance for all categories (no systematic bias)
- Predictions align with expert dermatologist consensus
- Include bias assessment and mitigation:
- Regular auditing of performance disparities across skin tones
- Documentation of dataset composition by skin tone category
- Strategies for addressing underrepresentation in training data
- Transparency reporting on per-type performance metrics
- Continuous monitoring of real-world performance across diverse populations
Clinical Impact:
The Skin Tone Identification model serves multiple critical functions:
- Bias mitigation: Enables skin tone-aware AI models that maintain equitable performance across all populations
- Treatment personalization: Supports accurate dosing for phototherapy, laser procedures, and skin-tone-specific therapeutics
- Research equity: Ensures clinical trials include and stratify diverse skin tones for representative evidence
- Quality assurance: Validates that dermatological AI systems perform equitably across all skin tone categories
- Regulatory compliance: Demonstrates AI model validation across diverse populations as required by regulatory agencies
- Clinical workflow integration: Provides automated skin tone documentation for electronic health records
Skin 3D Reconstruction
Description
This method transforms 2D pixel coordinates from an standard 2D image into 3D world metric coordinates, enabling comprehensive and accurate spatial analysis of skin surfaces.
This method leverages 3D metric maps and camera calibration parameters to convert pixel coordinates into real-world measurements, accounting for depth variations and perspective distortion.
For any given pixel coordinate , the 3D metric world coordinates are computed using the following equation:
Where:
- is the camera intrinsic matrix.
- is the metric depth value at pixel .
By applying this mathematical transformation to every point of a target surface segmentation, the method allows for straightforward geometric analysis, including the calculation of area, perimeter, axes, volume, and depth.
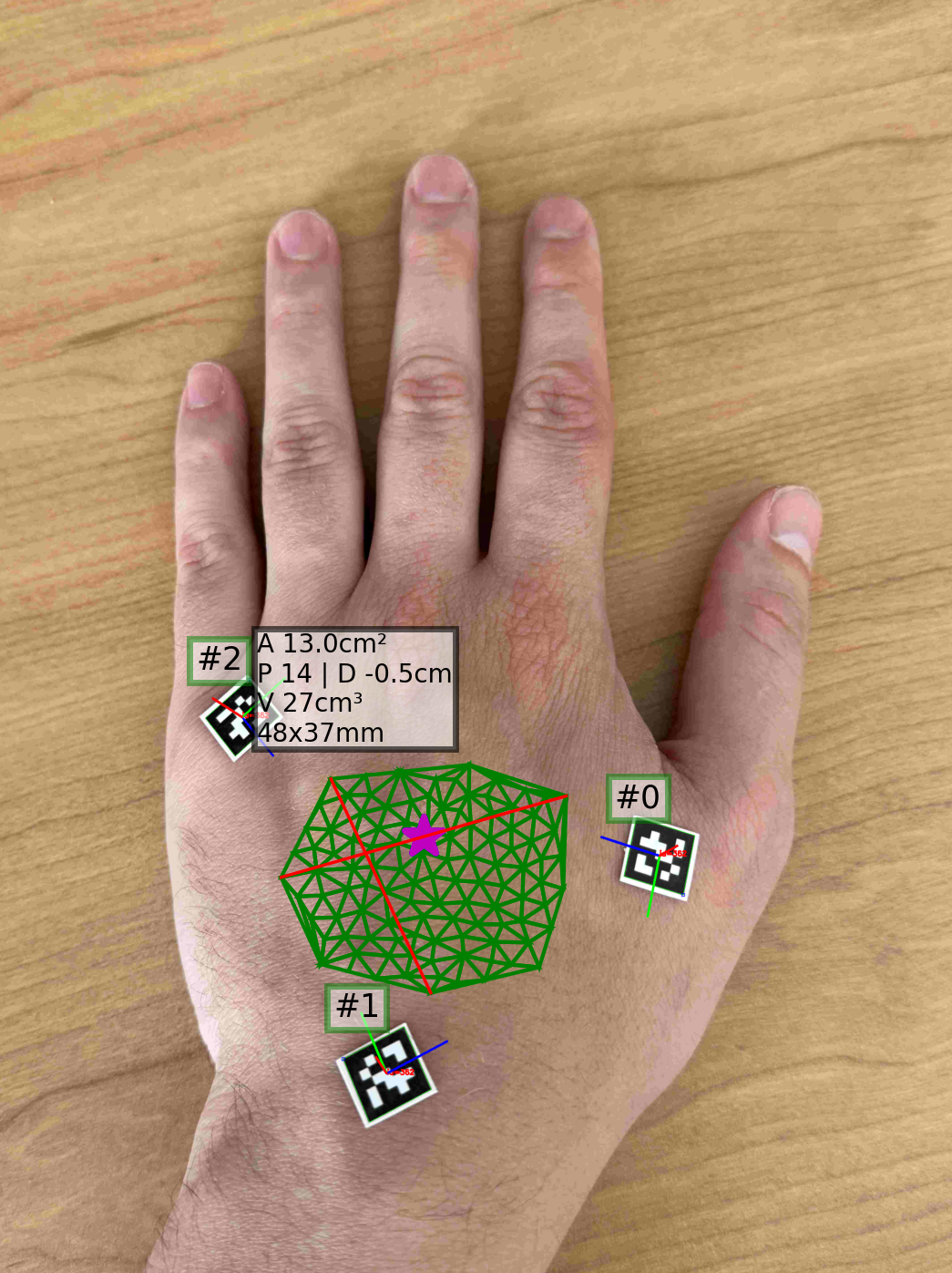 |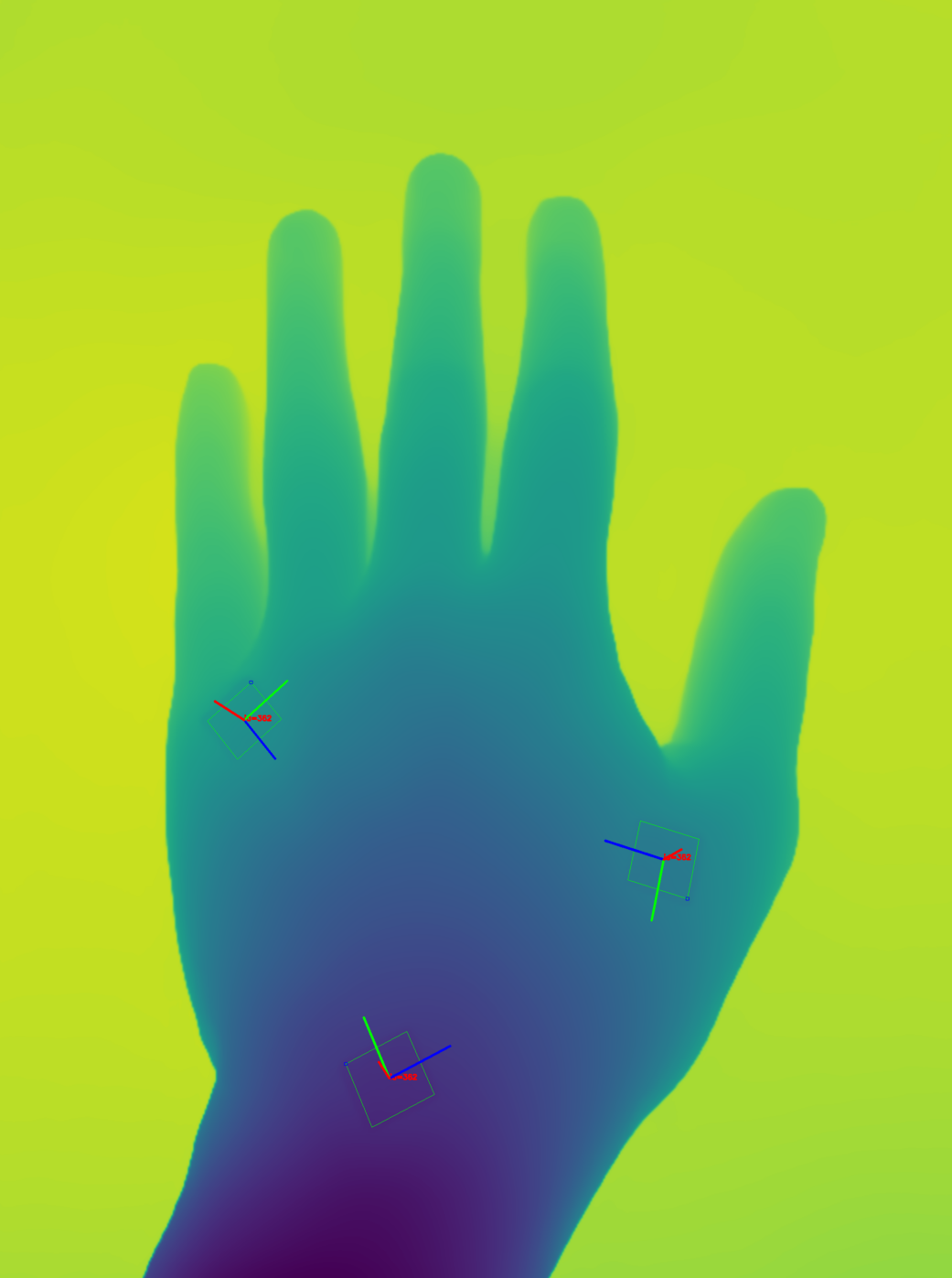  | 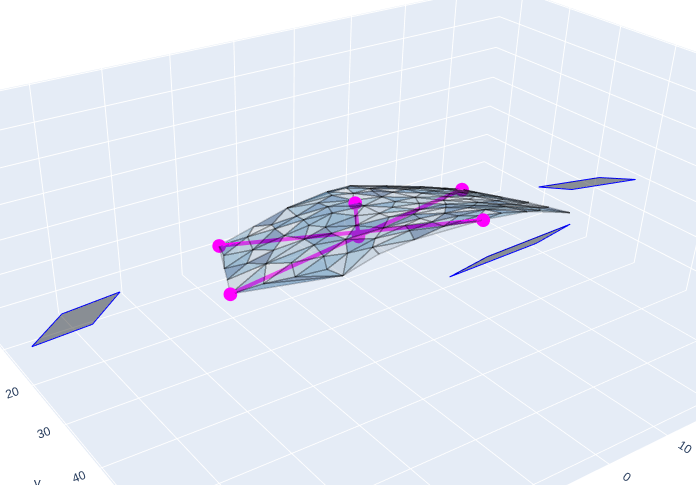 |
|---|
Left: Sample image with a delineated target surface and 3 reference markers. Results shows the area (A), perimeter (P), depth (D), volume (V), and dimensions (width x height) in cm. Center: depth map of the image and marker detections. Right: 3D visualization of the target surface, markers, and surface axes (width, height, and depth). Side perspective.
Objectives
- Enable accurate surface area quantification in body surface area (BSA) affected calculations for severity scoring systems (PASI, EASI, burn assessment, vitiligo VASI).
- Account for depth variation across non-planar body surfaces, providing more accurate measurements than simple 2D planimetry.
- Reduce measurement error associated with perspective distortion, camera angle, and irregular body surface curvature.
- Provide calibrated measurements in standardized physical units (e.g., mm²) for clinical documentation and research.
- Enable automated BSA percentage calculation by combining surface area measurements with body site identification.
- Support telemedicine workflows where physical ruler measurements are impractical or unavailable.
Justification (Clinical Evidence):
- Body surface area quantification is fundamental to severity scoring in dermatology, with PASI, EASI, and burn assessment all requiring accurate BSA affected estimates [275, 276].
- Manual BSA estimation shows high inter-observer variability (coefficient of variation 20-40%), particularly for irregular lesions or when visual estimation methods are used [277, 278].
- Simple 2D planimetry without depth correction introduces systematic errors of 15-35% when measuring non-planar body surfaces due to perspective distortion and surface curvature [279].
- Reference-based calibration has been validated in wound measurement showing accuracy within 5-10% of gold-standard methods (water displacement, 3D scanning) [280, 281].
- Monocular depth estimation combined with calibration markers achieves mean absolute error <8% for surface area quantification on curved surfaces [282].
- Automated BSA quantification improves reproducibility in clinical trials, with standardized measurements showing 50-70% reduction in outcome variability compared to visual estimation [283].
- Depth-aware surface area calculation is particularly critical for body sites with significant curvature (joints, torso, scalp) where 2D approximations introduce substantial error [284].
Evaluation and Area Calculation:
We evaluate the precision of the 3D reconstruction by evaluating the calculated area of shapes of known dimensions we draw on skin surfaces. Each target surface is analyzed by first triangulating it into a mesh of non-overlapping planar triangles using Delaunay triangulation. The corners of each triangle are then transformed into 3D world coordinates using the pixel-to-world transformation, allowing for the straightforward calculation of their area in real-world units. The segmented target area is calculated as the sum of the areas of all individual triangles, providing an accurate measurement that accounts for depth and curvature.
The computed area is then compared against the expected real-world area. The evaluation metric used is the Relative Mean Absolute Error (rMAE), requiring a strict confidence threshold of .
Endpoints and Requirements
Performance is evaluated using Relative Mean Absolute Error (rMAE) compared against the expected ground truth areas (known shapes drawn on the skin).
| Metric | Threshold | Interpretation |
|---|---|---|
| rMAE | ≤ 15% | Surface area estimates within 15% of relative ground truth area. |
All thresholds must be achieved with 95% confidence intervals.
Requirements:
- Implement a pixel to world coordinates transformation:
- Use precise depth information and camera intrinsics to transform pixel coordinates to 3D world coordinates.
- Demonstrate:
- Relative depth error ≤ 15% compared to ground truth depth measurements
- Consistent depth estimation across different body sites
- Implement surface triangulation
- Output structured data including:
- Total surface area (mm²) for each segmented region
- Perimeter of the region (mm)
- Maximum depth of the region (mm)
- Volume of the region (mm³)
- Axes lengths of the region (mm)
- Demonstrate:
- rMAE ≤ 15%
General Requirements
- Validate the full pipeline on an independent and diverse test dataset including:
- Various body sites: Face, neck, feet
- Different surface curvatures
- Various camera configurations: Distances, focal lengths, angles
- Ensure outputs are compatible with:
- PASI, EASI, VASI, burn assessment calculation systems
- Body surface area (BSA) estimation using Rule of Nines or Lund-Browder charts
- FHIR-based structured reporting for interoperability
- Clinical decision support systems requiring quantitative surface area data
- Research data collection for clinical trials
- Provide interpretability features:
- Visualization: Overlay of depth map, and segmented regions
- Document the pipeline architecture including:
- World information used as reference
- Depth estimation model architecture (pre-trained or custom)
- Integration strategy for multi-stage pipeline
- Provide evidence that:
- The pipeline generalizes across different body sites and curvatures
- Depth-aware calibration outperforms simple 2D planimetry
Clinical Impact:
The Surface Area Quantification pipeline serves critical functions:
- Accurate severity scoring: Enables precise BSA affected calculations for PASI, EASI, burn assessment, VASI
- Reproducible measurements: Reduces inter-observer variability in area estimation
- Depth-aware quantification: Accounts for body surface curvature
- Flexible deployment: Works with 1-N world reference information items
- Telemedicine enablement: Provides calibrated measurements from patient-captured images with reference information
- Clinical trial support: Standardizes surface area endpoints with objective, reproducible methodology
- Treatment monitoring: Enables accurate tracking of lesion size changes over time
Technical Details:
Reference World Information:
Reference world information might be necessary for precise real-world measurements. The requirements for this reference information might include:
- Detectable reference world information in images (markers): they must be located out of the region of interest, ideally at varying depths.
- LiDAR or depth sensor: If available, it can provide direct metric depth measurements without the need for reference markers.
If using markers as reference, they must meet the following criteria:
- Use planar, non-deformable markers to ensure accurate dimensional calibration
- Select matte (non-reflective) surfaces to prevent specular reflections and glare
- Include a white border of minimum 2-3 mm around the marker to ensure proper contrast and facilitate automated detection
- Verify marker dimensions are appropriate for the camera distance (e.g., minimum 10 mm marker size at approximately 30 cm distance)
- Position markers scattered across the image to capture depth variation, ideally at different distances from the camera
- Maintain markers in a planar orientation without strong perspective distortion
- Ensure markers are fully visible and completely within the image frame (no partial occlusion)
- Verify markers are captured free of shadows, reflections, and glare
- Ensure the white border is clearly visible and unobstructed
- Confirm markers are not deformed or bent during imaging
- Avoid placing markers in areas with uneven lighting or background interference
Depth Estimation Approach:
Monocular depth estimation might be needed to estimate relative depth without requiring specialized hardware
Camera Calibration:
- Intrinsic parameters: Estimated from reference markers such as from a ChArUco board, assuming pinhole camera model
Failure Modes and Limitations:
- No reference information detected: Cannot provide calibrated measurements
- Extreme camera angles: Perspective distortion may exceed correction capabilities, quality warning issued
- Poor depth estimation: Textureless regions or unusual body positions may yield unreliable depth, affecting calibration accuracy
Lighting Conditions
- Capture images under consistent, uniform lighting to avoid shadows and harsh highlights
- Use diffuse light sources to minimize reflections on skin and markers (if existent)
- Avoid direct sunlight or bright spot lighting that creates high-contrast shadows
- Ensure adequate illumination across the entire region of interest and markers (if existent)
Image Quality Standards
- Maintain consistent focus across both the region of interest and markers (if existent)
- Capture images at sufficient resolution
- Discard images with significant shadows, reflections, or occlusions and retake with corrected lighting and positioning
Color Correction
Description
The Color Correction model is a preprocessing algorithm designed to standardize the color representation of clinical images, adjusting the image so its colors closely reflect reality regardless of the original lighting conditions. The method works by identifying one or more standardized markers within the image that contain a grid of known reference colors. Let represent the observed color values extracted from the detected marker grids, and represent the theoretically expected true color values for those specific grids, the algorithm learns a color transformation function that maps the observed colors to their expected counterparts:
This method can compute this transformation using either a single reference marker or a set of multiple different markers scattered across the image. Once the optimal transformation function is learned, it is applied globally to every pixel in the original image to generate the standardized, color-corrected output :
 |
|---|
Left: Original image taken with low cool light and displaying marker detections. Center: image after color correction. Right: three rows representing the averaged detected colors, averaged colors after color correction, and the expected reference colors respectively.
Objectives
Justification (Clinical Evidence):
Endpoints and Requirements
Performance is evaluated by taking a set of images under different light temperatures, each containing the reference markers, learning the color transformation, and evaluating the error of the mapping against the known expected colors.
The error metric used is the standard color difference metric (Delta E) (CIEDE2000) in the CIELAB color space to account for a perceptually meaningful measure of color accuracy. The success criterion is set as , which ensures the color difference is not significant to the human eye.
| Metric | Threshold | Interpretation |
|---|---|---|
| ≤ 5.0 | The color difference is not significant to the human eye, ensuring high clinical color fidelity. |
All thresholds must be achieved with 95% confidence intervals.
Requirements:
- Scan the raw input image to locate the predefined reference marker (or markers) using robust computer vision techniques, such as contour analysis or keypoint matching
- Extract the observed RGB color values from the specific grid regions once the marker is precisely localized
- Apply an outlier rejection step to actively ignore false positive markers or markers with strong artifacts (e.g., shadows, reflections, occlusions) that could skew the color transformation
- Mathematically map the extracted observed values to the known, hardcoded reference values using an optimization algorithm to solve for the transformation function
- Apply the calculated function to the entire image array to adjust the global color balance to match reality while strictly preserving the structural, spatial, and textural details of the skin and lesions
- The corrected outputs are compatible with downstream clinical AI models.
- Demostrate ≤ 5.0.
General Requirements
- Validate the model on an independent and diverse test dataset including:
- Various lighting conditions (e.g., cool, warm, neutral)
- Variable amount of markers
- Integrate seamlessly into the image preprocessing pipeline before downstream clinical models (e.g., erythema or hyperpigmentation quantification) execute.
- Operate reliably across various camera hardware, including standard smartphones and professional clinical digital cameras.
- Detect and flag images where the reference marker is entirely missing, severely occluded, or illuminated beyond the mathematical limits of correctability.
- Maintain the original image resolution and aspect ratio during the transformation process.
Clinical Impact:
- Enhances the human interpretability of images for clinical decision-making by ensuring colors in the image closely match real-life appearance.
- Ensures accurate color representation for clinical AI models that rely on color features (e.g., erythema quantification, hyperpigmentation analysis).
- Enables highly accurate tracking of disease progression and treatment response over time by eliminating perceived color shifts caused by different lighting environments or camera devices.
- Provides standardized, normalized inputs to downstream clinical models, directly improving their reliability and reducing output variability.
- Empowers patients to capture actionable images from home in uncontrolled lighting conditions, ensuring the resulting images presented to the clinician accurately reflect reality.
- Prevents automated camera white-balance and exposure algorithms from disproportionately skewing the appearance of diverse skin tones (Fitzpatrick I-VI), ensuring equitable and accurate assessment across all patient demographics.
Technical Details:
-
Lighting Conditions
- Capture images under consistent, uniform lighting to avoid shadows and harsh highlights
- Use diffuse light sources to minimize reflections on skin and markers
- Avoid direct sunlight or bright spot lighting that creates high-contrast shadows
- Ensure adequate illumination across the entire region of interest and markers
-
Marker Properties
- Use planar, non-deformable markers to ensure accurate dimensional calibration
- Select matte (non-reflective) surfaces to prevent specular reflections and glare
- Include a white border of minimum 2-3 mm around the marker to ensure proper contrast and facilitate automated detection
- Verify marker dimensions are appropriate for the camera distance (e.g., minimum 10 mm marker size at approximately 30 cm distance)
-
Marker Positioning
- Position markers scattered across the image, to allow for accurate color correction across the entire field of view
- Maintain markers in a planar orientation without strong perspective distortion
- Ensure markers are fully visible and completely within the image frame (no partial occlusion)
-
Marker Appearance in Image
- Verify markers are captured free of shadows, reflections, and glare
- Ensure the white border is clearly visible and unobstructed
- Confirm markers are not deformed or bent during imaging
- Avoid placing markers in areas with uneven lighting or background interference
-
Image Quality Standards
- Maintain consistent focus across both the region of interest and markers
- Capture images at sufficient resolution to resolve marker details
- Verify the full region of interest and all reference markers are visible in a single image
- Discard images with significant shadows, reflections, or occlusions and retake with corrected lighting and positioning
Body Site Identification
Description
A deep learning multi-label classification model ingests an image and outputs a probability (sigmoid of the logits) distribution across anatomical body site categories:
where each corresponds to the probability that the image contains skin from anatomical site , where .
The model classifies images into the following body site categories:
Head and Neck Region:
- Face: All facial areas excluding scalp
- Top of the head: Perspective including crown region
- Back of the head: Perspective including occipital region
- Scalp: hair-bearing region
- Ear
- Eye
- Nose
- Mouth
- Tongue
- Neck
Upper Extremities:
- Arm
- Armpit
- Hand
- Finger
- Hand nail
Trunk:
- Back
- Trunk, chest and abdomen
Lower Extremities:
- Leg
- Knee
- Foot
- Toe
- Foot nail
Anogenital Region:
- Genitals (groin, penis, vulva, anus)
- Buttock
Other Specialized Sites:
- Close-up image (body part not visible)
To predicte body sites, a thresholding approach is used (typically with ) on the output probabilities. To obtain a binary predictions per body site, the following equation is applied:
where ( ) denotes presence of body site , and denotes absence.
Objectives
- Enable anatomical context awareness for downstream clinical AI models that may have site-specific performance characteristics or require body site information for clinical interpretation.
- Support body surface area (BSA) calculations by identifying anatomical regions for standardized BSA estimation in severity scoring systems (PASI, EASI, burn assessment).
- Facilitate disease-specific analysis by routing images to body site-appropriate clinical models (e.g., palmoplantar-specific psoriasis assessment, facial acne analysis).
- Improve clinical documentation by automatically annotating images with anatomical location for structured medical records.
- Enable epidemiological analysis by tracking disease distribution across body sites for research and surveillance purposes.
- Support treatment planning by providing anatomical context relevant for therapy selection (e.g., facial vs. truncal treatments differ in formulation and potency).
- Enhance quality control by detecting anatomically inappropriate images for specific clinical workflows.
Justification (Clinical Evidence):
- Body site location is a critical clinical variable influencing disease presentation, differential diagnosis, treatment selection, and prognosis across dermatological conditions [262, 263].
- Manual anatomical site annotation is time-consuming and inconsistent, with variability particularly evident for boundary regions (e.g., wrist vs. forearm, neck vs. chest) [264].
- Automated body site identification has demonstrated high accuracy (>85%) in multi-class classification tasks across diverse dermatological imaging datasets [265, 266].
- Disease prevalence, morphology, and treatment response vary significantly by anatomical site:
- Psoriasis: Scalp, elbows, knees show different treatment responses than intertriginous areas [267]
- Acne: Facial acne requires different therapeutic approaches than truncal acne [268]
- Hidradenitis suppurativa: Predominantly affects axillary, inguinal, and perianal regions [269]
- Melanoma: Sun-exposed sites (face, arms) have different risk profiles than trunk or acral sites [270]
- Body site-specific AI models show 10-20% accuracy improvement compared to site-agnostic models for certain conditions [271].
- Accurate body site identification enables automated BSA calculation for severity indices (PASI, EASI), where site-specific weighting is required (e.g., head = 10%, trunk = 30%, upper extremities = 20%, lower extremities = 40%) [272].
- Treatment guidelines often specify site-specific recommendations for corticosteroid potency, formulation selection, and therapy duration [273].
- Clinical trials require body site documentation for subgroup analyses and to ensure representative distribution of lesions [274].
Endpoints and Requirements
Performance is evaluated using Binary Accuracy (BAcc) per body site class.
| Metric | Threshold | Interpretation |
|---|---|---|
| BAcc. per each body part | ≥ 0.70 | Strong discrimination across body site classes. |
All thresholds must be achieved with 95% confidence intervals.
Requirements:
- Implement a deep learning multi-label classification architecture optimized for anatomical feature recognition.
- Report all metrics with 95% confidence intervals.
- Demonstrate performance meeting.
- Validate the model on an independent and diverse test dataset including:
- Broad anatomical region (head/neck, upper extremity, trunk, lower extremity, anogenital)
- Multiple imaging perspectives (frontal, lateral, oblique views)
- Diverse patient populations (various ages, genders, body habitus, Fitzpatrick skin types)
- Different imaging contexts (clinical photography, patient self-captured, telemedicine)
- Challenging boundary cases (wrist/hand, ankle/foot, neck/chest transitions)
- Partially visible anatomy where full body site context is limited
- Handle specialized anatomical features:
- Intertriginous regions: Axillae, inframammary, inguinal folds, perianal, interdigital
- Acral sites: Palms, soles, nail apparatus
- Mucosal surfaces: Oral mucosa, labial regions
- Hair-bearing regions: Scalp, beard area, body hair distribution
- Ensure outputs are compatible with:
- Body surface area (BSA) calculation algorithms for severity scoring (PASI, EASI, burn assessment)
- Site-specific clinical models requiring anatomical routing
- FHIR-based structured reporting with standardized anatomical site codes (SNOMED CT, ICD-11)
- Clinical decision support systems providing site-specific treatment recommendations
- Medical record systems for automated anatomical documentation
- Epidemiological databases for disease surveillance and research
- Handle challenging scenarios:
- Multi-site flags when multiple body regions are visible
- Ambiguous boundaries: Wrist/hand, ankle/foot, neck/shoulder transitions
- Close-ups: Limited anatomical context (e.g., extreme close-up of skin without landmarks)
- Provide evidence that:
- The model generalizes across different imaging devices and conditions
- Performance is maintained across diverse patient populations and body habitus
- Anatomical classification is robust to skin conditions and lesions
- The model handles partial anatomy and cropped images appropriately
- Multi-site detection works when multiple body regions are present
Clinical Impact:
The Body Site Identification model serves multiple critical functions:
- Automated documentation: Eliminates manual anatomical site entry in medical records
- BSA calculation support: Enables accurate body surface area estimation for severity indices
- Site-specific routing: Directs images to optimal body site-specific AI models
- Treatment personalization: Supports site-appropriate therapy recommendations
- Quality assurance: Validates anatomical appropriateness for specific clinical workflows
- Research enablement: Facilitates epidemiological analysis and clinical trial stratification
- Workflow optimization: Reduces clinician time spent on anatomical annotation
Data Specifications
The development of the AI models requires the systematic collection and annotation of dermatological images through two complementary approaches:
Archive Data
- Sources: Public medical datasets and dermatological atlases and private healthcare institution archives under data transfer agreements.
- Scale: Target >100,000 curated images from multiple validated sources.
- Coverage: Comprehensive ICD-11 diagnostic categories; both clinical and dermoscopic images; multiple continents and institutions; Fitzpatrick skin types I-VI.
- Validation: Diagnoses confirmed by board-certified dermatologists, histopathological analysis, or consensus expert opinion.
- Protocol Reference: R-TF-028-003 Data Collection Instructions - Archive Data.
Custom Gathered Data
- Sources: Clinical validation studies of Legit.Health Plus and dedicated prospective observational data acquisition studies.
- Operators: Board-certified dermatologists or trained clinical staff under dermatologist supervision.
- Image Specifications: 3-5 high-resolution images per case (≥1024×1024 pixels; ≥2000×2000 for dermoscopy); JPEG/PNG format; clinical and dermoscopic modalities; no manufacturer restrictions for real-world generalizability.
- Metadata: Primary diagnosis (ICD-11), differential diagnoses, diagnostic confidence, histopathology when available; age, sex, Fitzpatrick phototype; anatomical location, lesion characteristics; imaging modality and device.
- Ethical Compliance: IRB/CEIm approval, written informed consent, full GDPR compliance with de-identification, secure encrypted transfer.
- Protocol Reference: R-TF-028-003 Data Collection Instructions - Custom Gathered Data.
Data Quality and Reference Standard
- Quality Assurance: Resolution thresholds (≥200×200 dermoscopic, ≥400×400 clinical); automated artifact detection; diagnostic specificity validation; EXIF stripping; duplicate detection via perceptual hashing.
- Annotation: Exclusively by board-certified dermatologists; multi-expert consensus for ambiguous cases; histopathological confirmation when available; inter-rater reliability assessment (Cohen's kappa, ICC).
- Annotation Types (detailed in R-TF-028-004): ICD-11 diagnostic labels, clinical sign intensity, lesion segmentation, lesion counting, body site annotation, binary indicators.
Dataset Composition and Representativeness
The combined dataset reflects the intended use population:
- Demographics: Balanced Fitzpatrick skin types I-VI, age groups (pediatric to geriatric), sex distribution.
- Clinical Diversity: Hundreds of ICD-11 categories; mild to severe presentations; all major anatomical sites; clinical and dermoscopic modalities.
- Imaging Diversity: Multiple camera types (smartphones, clinical cameras, dermatoscopes); varied lighting, resolutions, framing; real-world artifacts.
Population characteristics and clinical representativeness documented in R-TF-028-005 AI Development Report.
Dataset Partitioning
- Training/Validation/Test Separation: Strict hold-out policies with patient-level splitting to prevent data leakage.
- Independence: Temporal and source separation where applicable; version control and comprehensive traceability.
Requirements:
- Execute retrospective and prospective data collection per R-TF-028-003 protocols.
- Ensure ethical compliance (IRB/CEIm approval, informed consent) and GDPR compliance.
- Implement rigorous quality assurance for images and labels.
- Establish robust reference standard via expert dermatologist annotation with inter-rater reliability assessment.
- Maintain strict test set independence from training/validation data.
- Document data provenance, demographic characteristics, and clinical representativeness in R-TF-028-005.
- Ensure demographic diversity across Fitzpatrick types, ages, and sex.
- Maintain version-controlled dataset management with change control procedures.
Other Specifications
Development Environment:
- Fixed hardware/software stack for training and evaluation.
- Deployment conversion validated by prediction equivalence testing.
Requirements:
- Track software versions (TensorFlow, NumPy, etc.).
- Verify equivalence between development and deployed model outputs.
Cybersecurity and Transparency
- Data: Always de-identified/pseudonymized [9].
- Access: Research server restricted to authorized staff only.
- Traceability: Development Report to include data management, model training, evaluation methods, and results.
- Explainability: Logs, saliency maps, and learning curves to support monitoring.
- User Documentation: Must state algorithm purpose, inputs/outputs, limitations, and that AI/ML is used.
Requirements:
- Secure and segregate research data.
- Provide full traceability of data and algorithms.
- Communicate limitations clearly to end-users.
Specifications and Risks
Risks linked to specifications are recorded in the AI/ML Risk Matrix (R-TF-028-011).
Key Risks:
- Misinterpretation of outputs.
- Incorrect diagnosis suggestions.
- Data bias or mislabeled reference standard.
- Model drift over time.
- Input image variability (lighting, resolution).
Risk Mitigations:
- Rigorous pre-market validation.
- Continuous monitoring and retraining.
- Controlled input requirements.
- Clear clinical instructions for use.
Integration and Environment
Integration
Algorithms will be packaged for integration into Legit.Health Plus to support healthcare professionals [20, 22, 25, 35].
Environment
- Inputs: Clinical and dermoscopic images [26].
- Robustness: Must handle variability in acquisition [8].
- Compatibility: Package size and computational load must align with target device hardware/software.
References
- Esteva A, Kuprel B, Novoa RA, et al. Dermatologist-level classification of skin cancer with deep neural networks. Nature. 2017;542(7639):115-118. doi:10.1038/nature21056
- Liu Y, Jain A, Eng C, et al. A deep learning system for differential diagnosis of skin diseases. Nat Med. 2020;26(6):900-908. doi:10.1038/s41591-020-0842-3
- Han SS, Kim MS, Lim W, Park GH, Park I, Chang SE. Classification of the clinical images for benign and malignant cutaneous tumors using a deep learning algorithm. J Invest Dermatol. 2018;138(7):1529-1538. doi:10.1016/j.jid.2018.01.028
- Haenssle HA, Fink C, Schneiderbauer R, et al. Man against machine: diagnostic performance of a deep learning convolutional neural network for dermoscopic melanoma recognition in comparison to 58 dermatologists. Ann Oncol. 2018;29(8):1836-1842. doi:10.1093/annonc/mdy166
- Brinker TJ, Hekler A, Enk AH, et al. Deep learning outperformed 136 of 157 dermatologists in a head-to-head dermoscopic melanoma image classification task. Eur J Cancer. 2019;113:47-54. doi:10.1016/j.ejca.2019.04.001
- Tschandl P, Codella N, Akay BN, et al. Comparison of the accuracy of human readers versus machine-learning algorithms for pigmented skin lesion classification: an open, web-based, international, diagnostic study. Lancet Oncol. 2019;20(7):938-947. doi:10.1016/S1470-2045(19)30333-X
- Selvaraju RR, Cogswell M, Das A, Vedantam R, Parikh D, Batra D. Grad-CAM: Visual explanations from deep networks via gradient-based localization. Int J Comput Vis. 2020;128(2):336-359. doi:10.1007/s11263-019-01228-7
- Janda M, Horsham C, Vagenas D, et al. Accuracy of mobile digital teledermoscopy for skin self-examinations in adults at high risk of skin cancer: an open-label, randomised controlled trial. Lancet Digit Health. 2020;2(3):e129-e137. doi:10.1016/S2589-7500(20)30001-7
- Han SS, Park I, Chang SE, et al. Augmented intelligence dermatology: deep neural networks empower medical professionals in diagnosing skin cancer and predicting treatment options for 134 skin disorders. J Invest Dermatol. 2020;140(9):1753-1761. doi:10.1016/j.jid.2020.01.019
- Rajpara SM, Botello AP, Townend J, Ormerod AD. Systematic review of dermoscopy and digital dermoscopy/artificial intelligence for the diagnosis of melanoma. Br J Dermatol. 2009;161(3):591-604. doi:10.1111/j.1365-2133.2009.09093.x
- Maron RC, Weichenthal M, Utikal JS, et al. Systematic outperformance of 112 dermatologists in multiclass skin cancer image classification by convolutional neural networks. Eur J Cancer. 2019;119:57-65. doi:10.1016/j.ejca.2019.06.013
- Tognetti L, Bonechi S, Andreini P, et al. A new deep learning approach integrated with clinical data for the dermoscopic differentiation of early melanomas from atypical nevi. J Dermatol Sci. 2021;101(2):115-122. doi:10.1016/j.jdermsci.2020.11.009
- Ferrante di Ruffano L, Dinnes J, Deeks JJ, et al. Optical coherence tomography for diagnosing skin cancer in adults. Cochrane Database Syst Rev. 2018;12(12):CD013189. doi:10.1002/14651858.CD013189
- Dinnes J, Deeks JJ, Chuchu N, et al. Dermoscopy, with and without visual inspection, for diagnosing melanoma in adults. Cochrane Database Syst Rev. 2018;12(12):CD011902. doi:10.1002/14651858.CD011902.pub2
- Phillips M, Marsden H, Jaffe W, et al. Assessment of accuracy of an artificial intelligence algorithm to detect melanoma in images of skin lesions. JAMA Netw Open. 2019;2(10):e1913436. doi:10.1001/jamanetworkopen.2019.13436
- National Institute for Health and Care Excellence (NICE). Suspected cancer: recognition and referral [NG12]. London: NICE; 2015. Updated 2021. Available from: https://www.nice.org.uk/guidance/ng12
- Garbe C, Amaral T, Peris K, et al. European consensus-based interdisciplinary guideline for melanoma. Part 1: Diagnostics - Update 2019. Eur J Cancer. 2020;126:141-158. doi:10.1016/j.ejca.2019.11.014
- Walter FM, Morris HC, Humphrys E, et al. Effect of adding a diagnostic aid to best practice to manage suspicious pigmented lesions in primary care: randomised controlled trial. BMJ. 2012;345:e4110. doi:10.1136/bmj.e4110
- Curchin DJ, Harris VR, McCormack CJ, et al. Early detection of melanoma: a consensus report from the Australian Skin and Skin Cancer Research Centre Melanoma Screening Summit. Aust J Gen Pract. 2022;51(1-2):9-14. doi:10.31128/AJGP-06-21-6016
- Warshaw EM, Gravely AA, Nelson DB. Reliability of physical examination versus lesion photography in assessing melanocytic skin lesion morphology. J Am Acad Dermatol. 2010;63(4):e81-e87. doi:10.1016/j.jaad.2009.11.030
- Tan J, Liu H, Leyden JJ, Leoni MJ. Reliability of clinician erythema assessment grading scale. J Am Acad Dermatol. 2014;71(4):760-763. doi:10.1016/j.jaad.2014.05.044
- Lee JH, Kim YJ, Kim J, et al. Erythema detection in digital skin images using CNN. Skin Res Technol. 2021;27(3):295-301. doi:10.1111/srt.12938
- Cho SB, Lee SJ, Chung WS, et al. Automated erythema detection and quantification in rosacea using deep learning. J Eur Acad Dermatol Venereol. 2021;35(4):965-972. doi:10.1111/jdv.17000
- Kim YJ, Park SH, Lee JH, et al. Automated erythema assessment using deep learning for sunscreen efficacy testing. Photodermatol Photoimmunol Photomed. 2023;39(2):135-142. doi:10.1111/phpp.12825
- Fredriksson T, Pettersson U. Severe psoriasis--oral therapy with a new retinoid. Dermatologica. 1978;157(4):238-244. doi:10.1159/000250839
- Langley RGB, Krueger GG, Griffiths CEM. Psoriasis: epidemiology, clinical features, and quality of life. Ann Rheum Dis. 2005;64(Suppl 2):ii18-ii23. doi:10.1136/ard.2004.033217
- Shen X, Zhang J, Yan C, Zhou H. An automatic diagnosis method of facial acne vulgaris based on convolutional neural network. Sci Rep. 2018;8(1):5839. doi:10.1038/s41598-018-24204-6
- Seité S, Khammari A, Benzaquen M, et al. Development and accuracy of an artificial intelligence algorithm for acne grading from smartphone photographs. Exp Dermatol. 2019;28(11):1252-1257. doi:10.1111/exd.14022
- Wu X, Wen N, Liang J, et al. Joint acne image grading and counting via label distribution learning. In: Proceedings of the IEEE/CVF International Conference on Computer Vision. 2019:10642-10651. doi:10.1109/ICCV.2019.01074
- Kimball AB, Kerdel F, Adams D, et al. Adalimumab for the treatment of moderate to severe hidradenitis suppurativa: a parallel randomized trial. Ann Intern Med. 2012;157(12):846-855. doi:10.7326/0003-4819-157-12-201212180-00004
- Olsen EA, Hordinsky MK, Price VH, et al. Alopecia areata investigational assessment guidelines--Part II. National Alopecia Areata Foundation. J Am Acad Dermatol. 2004;51(3):440-447. doi:10.1016/j.jaad.2003.09.032
- Lee Y, Lee SH, Kim YH, et al. Hair loss quantification from standardized scalp photographs using deep learning. J Invest Dermatol. 2022;142(6):1636-1643. doi:10.1016/j.jid.2021.10.031
- Messenger AG, McKillop J, Farrant P, McDonagh AJ, Sladden M. British Association of Dermatologists' guidelines for the management of alopecia areata 2012. Br J Dermatol. 2012;166(5):916-926. doi:10.1111/j.1365-2133.2012.10955.x
- Gardner SE, Frantz RA. Wound bioburden and infection-related complications in diabetic foot ulcers. Biol Res Nurs. 2008;10(1):44-53. doi:10.1177/1099800408319056
- Cutting KF, White RJ. Criteria for identifying wound infection--revisited. Ostomy Wound Manage. 2005;51(1):28-34.
- Rahma ON, Iyer R, Kattapuram T, et al. Objective assessment of perilesional erythema of chronic wounds using digital color image processing. Adv Skin Wound Care. 2015;28(1):11-16. doi:10.1097/01.ASW.0000459039.98700.74
- Schmitt J, Spuls PI, Thomas KS, et al. The Harmonising Outcome Measures for Eczema (HOME) statement to assess clinical signs of atopic eczema in trials. J Allergy Clin Immunol. 2014;134(4):800-807. doi:10.1016/j.jaci.2014.07.043
- Vakharia PP, Chopra R, Sacotte R, et al. Validation of patient-reported global severity of atopic dermatitis in adults. Allergy. 2018;73(2):451-458. doi:10.1111/all.13309
- Spuls PI, Lecluse LL, Poulsen ML, Bos JD, Stern RS, Nijsten T. How good are clinical severity and outcome measures for psoriasis?: quantitative evaluation in a systematic review. J Invest Dermatol. 2010;130(4):933-943. doi:10.1038/jid.2009.391
- Nast A, Jacobs A, Rosumeck S, Werner RN. Efficacy and safety of systemic long-term treatments for moderate-to-severe psoriasis: a systematic review and meta-analysis. J Invest Dermatol. 2015;135(11):2641-2648. doi:10.1038/jid.2015.206
- Esteva A, Chou K, Yeung S, et al. Deep learning-enabled medical computer vision. NPJ Digit Med. 2021;4(1):5. doi:10.1038/s41746-020-00376-2
- Thomsen K, Iversen L, Titlestad TL, Winther O. Systematic review of machine learning for diagnosis and prognosis in dermatology. J Dermatolog Treat. 2020;31(5):496-510. doi:10.1080/09546634.2019.1682500
- Młynek A, Zalewska-Janowska A, Martus P, Staubach P, Zuberbier T, Maurer M. How to assess disease activity in patients with chronic urticaria? Allergy. 2008;63(6):777-780. doi:10.1111/j.1398-9995.2008.01726.x
- Kinyanjui NM, Odongo T, Cintas C, et al. Estimating skin tone and effects on classification performance in dermatology datasets. arXiv preprint arXiv:1910.13268. 2019.
- Ezzedine K, Eleftheriadou V, Whitton M, van Geel N. Vitiligo. Lancet. 2015;386(9988):74-84. doi:10.1016/S0140-6736(14)60763-7
- Taylor SC, Arsonnaud S, Czernielewski J. The Taylor hyperpigmentation scale: a new visual assessment tool for the evaluation of skin color and pigmentation. Cutis. 2005;76(4):270-274.
- Del Bino S, Duval C, Bernerd F. Clinical and biological characterization of skin pigmentation diversity and its consequences on UV impact. Int J Mol Sci. 2018;19(9):2668. doi:10.3390/ijms19092668
- Ly BCK, Dyer EB, Feig JL, Chien AL, Del Bino S. Research Techniques Made Simple: Cutaneous Colorimetry: A Reliable Technique for Objective Skin Color Measurement. J Invest Dermatol. 2020;140(1):3-12.e1. doi:10.1016/j.jid.2019.11.003
- Tschandl P, Rosendahl C, Kittler H. The HAM10000 dataset, a large collection of multi-source dermatoscopic images of common pigmented skin lesions. Sci Data. 2018;5:180161. doi:10.1038/sdata.2018.161
- Winkler JK, Fink C, Toberer F, et al. Association between surgical skin markings in dermoscopic images and diagnostic performance of a deep learning convolutional neural network for melanoma recognition. JAMA Dermatol. 2019;155(10):1135-1141. doi:10.1001/jamadermatol.2019.1735
- Codella NCF, Gutman D, Celebi ME, et al. Skin lesion analysis toward melanoma detection: A challenge at the 2017 International symposium on biomedical imaging (ISBI), hosted by the international skin imaging collaboration (ISIC). In: 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018). IEEE; 2018:168-172. doi:10.1109/ISBI.2018.8363547
- Fujisawa Y, Inoue S, Nakamura Y. The possibility of deep learning-based, computer-aided skin tumor classifiers. Front Med (Lausanne). 2019;6:191. doi:10.3389/fmed.2019.00191
- Jain A, Way D, Gupta V, et al. Development and assessment of an artificial intelligence-based tool for skin condition diagnosis by primary care physicians and nurse practitioners in teledermatology practices. JAMA Netw Open. 2021;4(4):e217249. doi:10.1001/jamanetworkopen.2021.7249
- Lee T, Ng V, Gallagher R, Coldman A, McLean D. Dullrazor: A software approach to hair removal from images. Comput Biol Med. 1997;27(6):533-543. doi:10.1016/s0010-4825(97)00020-6
- Winkler JK, Sies K, Fink C, et al. Melanoma recognition by a deep learning convolutional neural network-Performance in different melanoma subtypes and localisations. Eur J Cancer. 2020;127:21-29. doi:10.1016/j.ejca.2019.11.020
- Chakravorty R, Abedini M, Halpern A, et al. Dermoscopic image segmentation using deep convolutional networks. In: 2017 39th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC). IEEE; 2017:542-545. doi:10.1109/EMBC.2017.8036895
- Bi L, Kim J, Ahn E, Kumar A, Fulham M, Feng D. Dermoscopic image segmentation via multistage fully convolutional networks. IEEE Trans Biomed Eng. 2017;64(9):2065-2074. doi:10.1109/TBME.2017.2712771
- Celebi ME, Wen Q, Iyatomi H, Shimizu K, Zhou H, Schaefer G. A state-of-the-art survey on lesion border detection in dermoscopy images. Dermoscopy Image Analysis. 2015:97-129. doi:10.1201/b19107-5
- Yuan Y, Chao M, Lo YC. Automatic skin lesion segmentation using deep fully convolutional networks with Jaccard distance. IEEE Trans Med Imaging. 2017;36(9):1876-1886. doi:10.1109/TMI.2017.2695227
- Mirikharaji Z, Abhishek K, Izadi S, Hamarneh G. Star shape prior in fully convolutional networks for skin lesion segmentation. In: Medical Image Computing and Computer Assisted Intervention – MICCAI 2018. Springer; 2018:737-745. doi:10.1007/978-3-030-00937-3_84
- Barata C, Ruela M, Francisco M, Mendonça T, Marques JS. Two systems for the detection of melanomas in dermoscopy images using texture and color features. IEEE Syst J. 2014;8(3):965-979. doi:10.1109/JSYST.2013.2271540
- Marchetti MA, Codella NCF, Dusza SW, et al. Results of the 2016 International Skin Imaging Collaboration International Symposium on Biomedical Imaging challenge: Comparison of the accuracy of computer algorithms to dermatologists for the diagnosis of melanoma from dermoscopic images. J Am Acad Dermatol. 2018;78(2):270-277.e1. doi:10.1016/j.jaad.2017.08.016
- Stern RS, Nijsten T, Feldman SR, Margolis DJ, Rolstad T. Psoriasis is common, carries a substantial burden even when not extensive, and is associated with widespread treatment dissatisfaction. J Investig Dermatol Symp Proc. 2004;9(2):136-139. doi:10.1046/j.1087-0024.2003.09102.x
- Jaspers S, Hopermann S, Sauermann G, et al. Rapid in vivo measurement of the topography of human skin by active image triangulation using a digital micromirror device. Skin Res Technol. 1999;5(3):195-207. doi:10.1111/j.1600-0846.1999.tb00131.x
- Takeshita J, Gelfand JM, Li P, et al. Psoriasis in the U.S. Medicare population: prevalence, treatment, and factors associated with biologic use. J Invest Dermatol. 2015;135(12):2955-2963. doi:10.1038/jid.2015.296
- Parisi R, Symmons DP, Griffiths CE, Ashcroft DM. Global epidemiology of psoriasis: a systematic review of incidence and prevalence. J Invest Dermatol. 2013;133(2):377-385. doi:10.1038/jid.2012.339
- Menter A, Strober BE, Kaplan DH, et al. Joint AAD-NPF guidelines of care for the management and treatment of psoriasis with biologics. J Am Acad Dermatol. 2019;80(4):1029-1072. doi:10.1016/j.jaad.2018.11.057
- Zaenglein AL, Pathy AL, Schlosser BJ, et al. Guidelines of care for the management of acne vulgaris. J Am Acad Dermatol. 2016;74(5):945-973.e33. doi:10.1016/j.jaad.2015.12.037
- Jemec GB. Clinical practice. Hidradenitis suppurativa. N Engl J Med. 2012;366(2):158-164. doi:10.1056/NEJMcp1014163
- Bradford PT, Goldstein AM, McMaster ML, Tucker MA. Acral lentiginous melanoma: incidence and survival patterns in the United States, 1986-2005. Arch Dermatol. 2009;145(4):427-434. doi:10.1001/archdermatol.2008.609
- Kawahara J, BenTaieb A, Hamarneh G. Deep features to classify skin lesions. In: 2016 IEEE 13th International Symposium on Biomedical Imaging (ISBI). IEEE; 2016:1397-1400. doi:10.1109/ISBI.2016.7493528
- Robinson A, Kardos M, Kimball AB. Physician Global Assessment (PGA) and Psoriasis Area and Severity Index (PASI): why do both? A systematic analysis of randomized controlled trials of biologic agents for moderate to severe plaque psoriasis. J Am Acad Dermatol. 2012;66(3):369-375. doi:10.1016/j.jaad.2011.01.022
- Yentzer BA, Ade RA, Fountain JM, et al. Simplifying regimens promotes greater adherence and outcomes with topical acne medications: a randomized controlled trial. Cutis. 2010;86(2):103-108.
- Rademaker M, Agnew K, Anagnostou N, et al. Psoriasis in those planning a family, pregnant or breastfeeding. The Australasian Psoriasis Collaboration. Australas J Dermatol. 2018;59(2):86-100. doi:10.1111/ajd.12733
- Brown BC, McKenna SP, Siddhi K, McGrouther DA, Bayat A. The hidden cost of skin scars: quality of life after skin scarring. J Plast Reconstr Aesthet Surg. 2008;61(9):1049-1058. doi:10.1016/j.bjps.2008.03.020
- Rennekampff HO, Hansbrough JF, Kiessig V, Doré C, Stoutenbeek CP, Schröder-Printzen I. Bioactive interleukin-8 is expressed in wounds and enhances wound healing. J Surg Res. 2000;93(1):41-54. doi:10.1006/jsre.2000.5892
- Wachtel TL, Berry CC, Wachtel EE, Frank HA. The inter-rater reliability of estimating the size of burns from various burn area chart drawings. Burns. 2000;26(2):156-170. doi:10.1016/s0305-4179(99)00047-9
- van Baar ME, Essink-Bot ML, Oen IM, Dokter J, Boxma H, van Beeck EF. Functional outcome after burns: a review. Burns. 2006;32(1):1-9. doi:10.1016/j.burns.2005.08.007
- Shuster S, Black MM, McVitie E. The influence of age and sex on skin thickness, skin collagen and density. Br J Dermatol. 1975;93(6):639-643. doi:10.1111/j.1365-2133.1975.tb05113.x
- Lucas C, Stanborough RW, Freeman CL, De Haan RJ. Efficacy of low-level laser therapy on wound healing in human subjects: a systematic review. Lasers Med Sci. 2000;15(2):84-93. doi:10.1007/s101030050053
- Mayrovitz HN, Soontupe LB. Wound areas by computerized planimetry of digital images: accuracy and reliability. Adv Skin Wound Care. 2009;22(5):222-229. doi:10.1097/01.ASW.0000305410.58350.36
- Wannous H, Lucas Y, Treuillet S, Albouy B. Supervised tissue classification from color images for a complete wound assessment tool. In: 2007 29th Annual International Conference of the IEEE Engineering in Medicine and Biology Society. IEEE; 2007:6031-6034. doi:10.1109/IEMBS.2007.4353725
- Spilsbury K, Semmens JB, Saunders CM, Hall SE. Long-term survival outcomes following breast-conserving surgery with and without radiotherapy for invasive breast cancer. ANZ J Surg. 2005;75(5):337-342. doi:10.1111/j.1445-2197.2005.03374.x
- Draaijers LJ, Tempelman FR, Botman YA, et al. The patient and observer scar assessment scale: a reliable and feasible tool for scar evaluation. Plast Reconstr Surg. 2004;113(7):1960-1965. doi:10.1097/01.prs.0000122207.28773.56
- Lebwohl M, Yeilding N, Szapary P, et al. Impact of weight on the efficacy and safety of ustekinumab in patients with moderate to severe psoriasis: rationale for dosing recommendations. J Am Acad Dermatol. 2010;63(4):571-579. doi:10.1016/j.jaad.2009.11.012
- Hanifin JM, Thurston M, Omoto M, Cherill R, Tofte SJ, Graeber M. The eczema area and severity index (EASI): assessment of reliability in atopic dermatitis. EASI Evaluator Group. Exp Dermatol. 2001;10(1):11-18. doi:10.1034/j.1600-0625.2001.100102.x
- Thomas CL, Finlay KA. Defining the boundaries: a critical evaluation of the Birmingham Burn Unit body map. Burns. 1986;12(8):544-548. doi:10.1016/0305-4179(86)90188-1
- Berkley JL. Determining total body surface area of a burn using a Lund and Browder chart. Nursing. 2007;37(10):18. doi:10.1097/01.NURSE.0000296227.88874.9e
- Langley RG, Ellis CN. Evaluating psoriasis with Psoriasis Area and Severity Index, Psoriasis Global Assessment, and Lattice System Physician's Global Assessment. J Am Acad Dermatol. 2004;51(4):563-569. doi:10.1016/j.jaad.2004.04.012
- Finlay AY. Current severe psoriasis and the rule of tens. Br J Dermatol. 2005;152(5):861-867. doi:10.1111/j.1365-2133.2005.06502.x
- Gudi V, Akhondi H. Burn Surface Area Assessment. In: StatPearls. Treasure Island (FL): StatPearls Publishing; 2023.
- Hawro T, Ohanyan T, Schoepke N, et al. The urticaria activity score—validity, reliability, and responsiveness. J Allergy Clin Immunol Pract. 2018;6(4):1185-1190.e1. doi:10.1016/j.jacp.2017.10.001
- Maurer M, Weller K, Bindslev-Jensen C, et al. Unmet clinical needs in chronic spontaneous urticaria. A GA²LEN task force report. Allergy. 2011;66(3):317-330. doi:10.1111/j.1398-9995.2010.02496.x
- Han SS, Moon IJ, Lim W, et al. Keratinocytic skin cancer detection on the face using region-based convolutional neural network. JAMA Dermatol. 2020;156(1):29-37. doi:10.1001/jamadermatol.2019.3807
- Zuberbier T, Balke M, Worm M, Edenharter G, Maurer M. Epidemiology of urticaria: a representative cross-sectional population survey. Clin Exp Dermatol. 2010;35(8):869-873. doi:10.1111/j.1365-2230.2010.03840.x
- Mathias SD, Dreskin SC, Kaplan A, Saini SS, Rosén K, Beck LA. Development of a daily diary for patients with chronic idiopathic urticaria. Ann Allergy Asthma Immunol. 2010;105(2):142-148. doi:10.1016/j.anai.2010.06.011
- Olsen EA, Dunlap FE, Funicella T, et al. A randomized clinical trial of 5% topical minoxidil versus 2% topical minoxidil and placebo in the treatment of androgenetic alopecia in men. J Am Acad Dermatol. 2002;47(3):377-385. doi:10.1067/mjd.2002.124088
- Alkhalifah A, Alsantali A, Wang E, McElwee KJ, Shapiro J. Alopecia areata update: part I. Clinical picture, histopathology, and pathogenesis. J Am Acad Dermatol. 2010;62(2):177-188. doi:10.1016/j.jaad.2009.10.032
- Rich P, Scher RK. Nail Psoriasis Severity Index: a useful tool for evaluation of nail psoriasis. J Am Acad Dermatol. 2003;49(2):206-212. doi:10.1067/s0190-9622(03)00910-1
- Fernández-Nieto D, Cura-Gonzalez ID, Esteban-Velasco C, Marques-Mejias MA, Ortega-Quijano D. Artificial intelligence to assess nail unit disorders: A pilot study. Skin Appendage Disord. 2021;7(6):428-433. doi:10.1159/000517341
- Parrish CA, Sobera JO, Elewski BE. Modification of the Nail Psoriasis Severity Index. J Am Acad Dermatol. 2005;53(4):745-746. doi:10.1016/j.jaad.2005.04.028
- Antonini D, Simonatto M, Candi E, Melino G. Keratinocyte stem cells and their niches in the skin appendages. J Invest Dermatol. 2014;134(7):1797-1799. doi:10.1038/jid.2014.126
- Njoo MD, Westerhof W, Bos JD, Bossuyt PM. A systematic review of autologous transplantation methods in vitiligo. Arch Dermatol. 1998;134(12):1543-1549. doi:10.1001/archderm.134.12.1543
- Grimes PE, Miller MM. Vitiligo: Patient stories, self-esteem, and the psychological burden of disease. Int J Womens Dermatol. 2018;4(1):32-37. doi:10.1016/j.ijwd.2017.11.005
- Hamzavi I, Jain H, McLean D, Shapiro J, Zeng H, Lui H. Parametric modeling of narrowband UV-B phototherapy for vitiligo using a novel quantitative tool: the Vitiligo Area Scoring Index. Arch Dermatol. 2004;140(6):677-683. doi:10.1001/archderm.140.6.677
- Njoo MD, Vodegel RM, Westerhof W. Depigmentation therapy in vitiligo universalis with topical 4-methoxyphenol and the Q-switched ruby laser. J Am Acad Dermatol. 2000;42(5 Pt 1):760-769. doi:10.1016/s0190-9622(00)90009-x
- Parsad D, Pandhi R, Dogra S, Kumar B. Clinical study of repigmentation patterns with different treatment modalities and their correlation with speed and stability of repigmentation in 352 vitiliginous patches. J Am Acad Dermatol. 2004;50(1):63-67. doi:10.1016/s0190-9622(03)02463-4
- Rodrigues M, Ezzedine K, Hamzavi I, Pandya AG, Harris JE. New discoveries in the pathogenesis and classification of vitiligo. J Am Acad Dermatol. 2017;77(1):1-13. doi:10.1016/j.jaad.2016.10.048
- Passeron T, Ortonne JP. Use of the 308-nm excimer laser for psoriasis and vitiligo. Clin Dermatol. 2006;24(1):33-42. doi:10.1016/j.clindermatol.2005.10.018
- Gawkrodger DJ, Ormerod AD, Shaw L, et al. Guideline for the diagnosis and management of vitiligo. Br J Dermatol. 2008;159(5):1051-1076. doi:10.1111/j.1365-2133.2008.08881.x
- Halder RM, Taliaferro SJ. Vitiligo. In: Wolff K, Goldsmith LA, Katz SI, et al, eds. Fitzpatrick's Dermatology in General Medicine. 7th ed. McGraw-Hill; 2008:616-622.
- Tan JK, Tang J, Fung K, et al. Development and validation of a comprehensive acne severity scale. J Cutan Med Surg. 2007;11(6):211-216. doi:10.2310/7750.2007.00037
- Layton AM, Henderson CA, Cunliffe WJ. A clinical evaluation of acne scarring and its incidence. Clin Exp Dermatol. 1994;19(4):303-308. doi:10.1111/j.1365-2230.1994.tb01200.x
- Tan J, Thiboutot D, Popp G, et al. Randomized phase 3 evaluation of trifarotene 50 μg/g cream treatment of moderate facial and truncal acne. J Am Acad Dermatol. 2019;80(6):1691-1699. doi:10.1016/j.jaad.2019.02.044
- Leyden J, Stein-Gold L, Weiss J. Why topical retinoids are mainstay of therapy for acne. Dermatol Ther (Heidelb). 2017;7(3):293-304. doi:10.1007/s13555-017-0185-2
- Thiboutot DM, Dréno B, Abanmi A, et al. Practical management of acne for clinicians: An international consensus from the Global Alliance to Improve Outcomes in Acne. J Am Acad Dermatol. 2018;78(2 Suppl 1):S1-S23.e1. doi:10.1016/j.jaad.2017.09.078
- Seité S, Dréno B, Benech F, Bédane C, Pecastaings S. Creation and validation of an artificial intelligence algorithm for acne grading. J Eur Acad Dermatol Venereol. 2020;34(12):2946-2951. doi:10.1111/jdv.16736
- Winkler JK, Sies K, Fink C, et al. Association between different scale bars in dermoscopic images and diagnostic performance of a market-approved deep learning convolutional neural network for melanoma recognition. Eur J Cancer. 2021;145:146-154. doi:10.1016/j.ejca.2020.12.010
- Burlina P, Joshi N, Ng E, Billings S, Paul W, Rotemberg V. Assessment of deep generative models for high-resolution synthetic retinal image generation of age-related macular degeneration. JAMA Ophthalmol. 2019;137(3):258-264. doi:10.1001/jamaophthalmol.2018.6156
- Korotkov K, Garcia R. Computerized analysis of pigmented skin lesions: A review. Artif Intell Med. 2012;56(2):69-90. doi:10.1016/j.artmed.2012.08.002
- Brinker TJ, Hekler A, Hauschild A, et al. Comparing artificial intelligence algorithms to 157 German dermatologists: the melanoma classification benchmark. Eur J Cancer. 2019;111:30-37. doi:10.1016/j.ejca.2018.12.016
- Perednia DA, Brown NA. Teledermatology: one application of telemedicine. Bull Med Libr Assoc. 1995;83(1):42-47.
- Ngoo A, Finnane A, McMeniman E, Tan JM, Janda M, Soyer HP. Fighting melanoma with smartphones: A snapshot on where we are a decade after app stores opened their doors. Int J Med Inform. 2018;118:99-112. doi:10.1016/j.ijmedinf.2018.08.004
- Kroemer S, Frühauf J, Campbell TM, et al. Mobile teledermatology for skin tumour screening: diagnostic accuracy of clinical and dermoscopic image tele-evaluation using cellular phones. Br J Dermatol. 2011;164(5):973-979. doi:10.1111/j.1365-2133.2011.10208.x
- Massone C, Hofmann-Wellenhof R, Ahlgrimm-Siess V, Gabler G, Ebner C, Soyer HP. Melanoma screening with cellular phones. PLoS One. 2007;2(5):e483. doi:10.1371/journal.pone.0000483
- Ferrara G, Argenziano G, Soyer HP, et al. The influence of clinical information in the histopathologic diagnosis of melanocytic skin neoplasms. PLoS One. 2009;4(4):e5375. doi:10.1371/journal.pone.0005375
- Carli P, De Giorgi V, Crocetti E, et al. Improvement of malignant/benign ratio in excised melanocytic lesions in the 'dermoscopy era': a retrospective study 1997-2001. Br J Dermatol. 2004;150(4):687-692. doi:10.1111/j.0007-0963.2004.05860.x
- Del Bino S, Bernerd F. Variations in skin colour and the biological consequences of ultraviolet radiation exposure. Br J Dermatol. 2013;169(Suppl 3):33-40. doi:10.1111/bjd.12529
- Pershing S, Enns JT, Bae IS, Randall BD, Pruiksma JB, Desai AD. Variability in physician assessment of oculoplastic standardized photographs. Aesthet Surg J. 2014;34(8):1203-1209. doi:10.1177/1090820X14542642
- Goh CL. The need for evidence-based aesthetic dermatology practice. J Cutan Aesthet Surg. 2009;2(2):65-71. doi:10.4103/0974-2077.58518
- Lester JC, Jia JL, Zhang L, Okoye GA, Linos E. Absence of images of skin of colour in publications of COVID-19 skin manifestations. Br J Dermatol. 2020;183(3):593-595. doi:10.1111/bjd.19258
- Wagner JK, Jovel C, Norton HL, Parra EJ, Shriver MD. Comparing quantitative measures of erythema, pigmentation and skin response using reflectometry. Pigment Cell Res. 2002;15(5):379-384. doi:10.1034/j.1600-0749.2002.02042.x
- Nkengne A, Bertin C, Stamatas GN, et al. Influence of facial skin attributes on the perceived age of Caucasian women. J Eur Acad Dermatol Venereol. 2008;22(8):982-991. doi:10.1111/j.1468-3083.2008.02698.x
- Davenport T, Kalakota R. The potential for artificial intelligence in healthcare. Future Healthc J. 2019;6(2):94-98. doi:10.7861/futurehosp.6-2-94
- Char DS, Shah NH, Magnus D. Implementing machine learning in health care - addressing ethical challenges. N Engl J Med. 2018;378(11):981-983. doi:10.1056/NEJMp1714229
- Gichoya JW, Banerjee I, Bhimireddy AR, et al. AI recognition of patient race in medical imaging: a modelling study. Lancet Digit Health. 2022;4(6):e406-e414. doi:10.1016/S2589-7500(22)00063-2
- Alexis AF, Sergay AB, Taylor SC. Common dermatologic disorders in skin of color: a comparative practice survey. Cutis. 2007;80(5):387-394.
- Ebede TL, Arch EL, Berson D. Hormonal treatment of acne in women. J Clin Aesthet Dermatol. 2009;2(12):16-22.
- Ly BC, Dyer EB, Feig JL, Chien AL, Del Bino S. Research Techniques Made Simple: Cutaneous Colorimetry: A Reliable Technique for Objective Skin Color Measurement. J Invest Dermatol. 2020;140(1):3-12.e1. doi:10.1016/j.jid.2019.11.003
- Chardon A, Cretois I, Hourseau C. Skin colour typology and suntanning pathways. Int J Cosmet Sci. 1991;13(4):191-208. doi:10.1111/j.1467-2494.1991.tb00561.x
- Gareau DS. Feasibility of digitally stained multimodal confocal mosaics to simulate histopathology. J Biomed Opt. 2009;14(3):034050. doi:10.1117/1.3149853
- Koenig K, Raphael AP, Lin L, et al. Optical skin biopsies by clinical CARS and multiphoton fluorescence/SHG tomography. Laser Phys Lett. 2011;8(6):465-468. doi:10.1002/lapl.201110014
- Baldi A, Murace R, Dragonetti E, et al. The Significance of Artificial Intelligence in the Assessment of Skin Cancer. J Clin Med. 2021;10(21):4926. doi:10.3390/jcm10214926
- Esteva A, Robicquet A, Ramsundar B, et al. A guide to deep learning in healthcare. Nat Med. 2019;25(1):24-29. doi:10.1038/s41591-018-0316-z
- Winkler JK, Fink C, Toberer F, et al. Association between surgical skin markings in dermoscopic images and diagnostic performance of a deep learning convolutional neural network for melanoma recognition. JAMA Dermatol. 2019;155(10):1135-1141. doi:10.1001/jamadermatol.2019.1735
- Kittler H, Pehamberger H, Wolff K, Binder M. Diagnostic accuracy of dermoscopy. Lancet Oncol. 2002;3(3):159-165. doi:10.1016/s1470-2045(02)00679-4
- Beam AL, Kohane IS. Big data and machine learning in health care. JAMA. 2018;319(13):1317-1318. doi:10.1001/jama.2017.18391
- Codella NC, Lin CC, Halpern A, et al. Collaborative human-AI (CHAI): Evidence-based interpretable melanoma classification in dermoscopic images. In: Understanding and Interpreting Machine Learning in Medical Image Computing Applications. Springer; 2018:97-105. doi:10.1007/978-3-030-02628-8_11
- Zouboulis CC, Desai N, Emtestam L, et al. European S1 guideline for the treatment of hidradenitis suppurativa/acne inversa. J Eur Acad Dermatol Venereol. 2015;29(4):619-644. doi:10.1111/jdv.12966
- Martorell A, García-Martínez FJ, Jiménez-Gallo D, et al. An update on hidradenitis suppurativa (Part I): Epidemiology, clinical aspects, and definition of disease severity. Actas Dermosifiliogr. 2015;106(9):703-715. doi:10.1016/j.ad.2015.06.004
- Hurley HJ. Axillary hyperhidrosis, apocrine bromhidrosis, hidradenitis suppurativa, and familial benign pemphigus: surgical approach. In: Roenigk RK, Roenigk HH Jr, eds. Dermatologic Surgery: Principles and Practice. Marcel Dekker; 1989:623-645.
- Revuz JE, Canoui-Poitrine F, Wolkenstein P, et al. Prevalence and factors associated with hidradenitis suppurativa: results from two case-control studies. J Am Acad Dermatol. 2008;59(4):596-601. doi:10.1016/j.jaad.2008.06.020
- Alikhan A, Sayed C, Alavi A, et al. North American clinical management guidelines for hidradenitis suppurativa: A publication from the United States and Canadian Hidradenitis Suppurativa Foundations: Part I: Diagnosis, evaluation, and the use of complementary and procedural management. J Am Acad Dermatol. 2019;81(1):76-90. doi:10.1016/j.jaad.2019.02.067
- Ingram JR, Collier F, Brown D, et al. British Association of Dermatologists guidelines for the management of hidradenitis suppurativa (acne inversa) 2018. Br J Dermatol. 2019;180(5):1009-1017. doi:10.1111/bjd.17537
- Esmann S, Jemec GB. Psychosocial impact of hidradenitis suppurativa: a qualitative study. Acta Derm Venereol. 2011;91(3):328-332. doi:10.2340/00015555-1082
- Matusiak Ł, Bieniek A, Szepietowski JC. Increased serum tumour necrosis factor-α in hidradenitis suppurativa patients: is there a basis for treatment with anti-tumour necrosis factor-α agents? Acta Derm Venereol. 2009;89(6):601-603. doi:10.2340/00015555-0701
- Grant A, Gonzalez T, Montgomery MO, Cardenas V, Kerdel FA. Infliximab therapy for patients with moderate to severe hidradenitis suppurativa: a randomized, double-blind, placebo-controlled crossover trial. J Am Acad Dermatol. 2010;62(2):205-217. doi:10.1016/j.jaad.2009.06.050
- Schneider-Burrus S, Tsaousi A, Barbus S, Huss-Marp J, Witte-Händel E, Witte K. Features associated with quality of life impairment in hidradenitis suppurativa patients. Front Med (Lausanne). 2021;8:676241. doi:10.3389/fmed.2021.676241
- Moriarty B, Jiyad Z, Creamer D. Four-weekly infliximab in the treatment of severe hidradenitis suppurativa. Br J Dermatol. 2014;170(4):986-987. doi:10.1111/bjd.12823
- Vossen ARJV, van der Zee HH, Prens EP. Hidradenitis Suppurativa: A Systematic Review Integrating Inflammatory Pathways Into a Cohesive Pathogenic Model. Front Immunol. 2018;9:2965. doi:10.3389/fimmu.2018.02965
- Sabat R, Jemec GBE, Matusiak Ł, Kimball AB, Prens E, Wolk K. Hidradenitis suppurativa. Nat Rev Dis Primers. 2020;6(1):18. doi:10.1038/s41572-020-0149-1
- Kimball AB, Kerdel F, Adams D, et al. Adalimumab for the treatment of moderate to severe hidradenitis suppurativa: a parallel randomized trial. Ann Intern Med. 2012;157(12):846-855. doi:10.7326/0003-4819-157-12-201212180-00004
- Zouboulis CC, Tzellos T, Kyrgidis A, et al. Development and validation of the International Hidradenitis Suppurativa Severity Score System (IHS4), a novel dynamic scoring system to assess HS severity. Br J Dermatol. 2017;177(5):1401-1409. doi:10.1111/bjd.15748
- Kimball AB, Okun MM, Williams DA, et al. Two Phase 3 Trials of Adalimumab for Hidradenitis Suppurativa. N Engl J Med. 2016;375(5):422-434. doi:10.1056/NEJMoa1504370
- Jfri A, Nassim D, O'Brien E, Gulliver W, Nikolakis G, Zouboulis CC. Prevalence of Hidradenitis Suppurativa: A Systematic Review and Meta-regression Analysis. JAMA Dermatol. 2021;157(8):924-931. doi:10.1001/jamadermatol.2021.1677
- Gomolin A, Cline A, Russo S, Wirya SA, Treat JR. Treatment of inflammatory manifestations of hidradenitis suppurativa with secukinumab in pediatric patients. JAAD Case Rep. 2019;5(12):1088-1091. doi:10.1016/j.jdcr.2019.10.005
- Mehdizadeh A, Hazen PG, Bechara FG, et al. Recurrence of hidradenitis suppurativa after surgical management: A systematic review and meta-analysis. J Am Acad Dermatol. 2015;73(5 Suppl 1):S70-S77. doi:10.1016/j.jaad.2015.07.044
- Goyal M, Knackstedt T, Yan S, Hassanpour S. Artificial intelligence-based image classification methods for diagnosis of skin cancer: Challenges and opportunities. Comput Biol Med. 2020;127:104065. doi:10.1016/j.compbiomed.2020.104065
- Nasr-Esfahani E, Samavi S, Karimi N, et al. Melanoma detection by analysis of clinical images using convolutional neural network. Annu Int Conf IEEE Eng Med Biol Soc. 2016;2016:1373-1376. doi:10.1109/EMBC.2016.7590963
- Garnavi R, Aldeen M, Celebi ME, Varigos G, Finch S. Border detection in dermoscopy images using hybrid thresholding on optimized color channels. Comput Med Imaging Graph. 2011;35(2):105-115. doi:10.1016/j.compmedimag.2010.08.001
- Xie Y, Zhang J, Xia Y. Semi-supervised adversarial model for benign-malignant lung nodule classification on chest CT. Med Image Anal. 2019;57:237-248. doi:10.1016/j.media.2019.07.004
- Serte S, Serener A, Al-Turjman F. Deep learning in medical imaging: A brief review. Trans Emerg Telecommun Technol. 2020;e4080. doi:10.1002/ett.4080
- Lee H, Chen YP. Image based computer aided diagnosis system for cancer detection. Expert Syst Appl. 2015;42(12):5356-5365. doi:10.1016/j.eswa.2015.02.005
- Udrea A, Mitra GD, Costea D, et al. Accuracy of a smartphone application for triage of skin lesions based on machine learning algorithms. J Eur Acad Dermatol Venereol. 2020;34(3):648-655. doi:10.1111/jdv.15935
- Schmid-Saugeón P, Guillod J, Thiran JP. Towards a computer-aided diagnosis system for pigmented skin lesions. Comput Med Imaging Graph. 2003;27(1):65-78. doi:10.1016/s0895-6111(02)00048-4
- Kasmi R, Mokrani K. Classification of malignant melanoma and benign skin lesions: implementation of automatic ABCD rule. IET Image Process. 2016;10(6):448-455. doi:10.1049/iet-ipr.2015.0385
- Jemec GB, Heidenheim M, Nielsen NH. The prevalence of hidradenitis suppurativa and its potential precursor lesions. J Am Acad Dermatol. 1996;35(2 Pt 1):191-194. doi:10.1016/s0190-9622(96)90321-7
- Yanling LI, Adams Wai-Kin Kong, Steven Thng. Segmenting vitiligo on clinical face images using CNN trained on synthetic and internet images. IEEE Journal of Biomedical and Health Informatics. 2021;25(8):3082-3093. doi:10.1109/JBHI.2021.3063388
- Guo L, et al. A deep learning-based hybrid artificial intelligence model for the detection and severity assessment of vitiligo lesions. Ann Transl Med. 2022;10(10):590. doi:10.21037/atm-22-1738
- Lu J, et al. Automatic segmentation of scaling in 2-D psoriasis skin images. IEEE Trans Med Imaging. 2012;32(4):719-730. doi:10.1109/TMI.2012.2236349
- Hashemifard K, Florez-Revuelta F. From garment to skin: the visuaal skin segmentation dataset. InInternational conference on image analysis and processing 2022 May 23 (pp. 59-70). Cham: Springer International Publishing.
- Olsen EA, Hordinsky MK, Price VH, Roberts JL, Shapiro J, Canfield D, Duvic M, King LE, McMichael AJ, Randall VA, Turner ML. Alopecia areata investigational assessment guidelines–Part II. Journal of the American Academy of Dermatology. 2004 Sep 1;51(3):440-7.
- Dash, Manoranjan, et al. "PsLSNet: Automated psoriasis skin lesion segmentation using modified U-Net-based fully convolutional network." Biomedical Signal Processing and Control 52 (2019): 226-237.
- Raj, Ritesh, Narendra D. Londhe, and Rajendra Sonawane. "Automated psoriasis lesion segmentation from unconstrained environment using residual U-Net with transfer learning." Computer Methods and Programs in Biomedicine 206 (2021): 106123.
- Scebba, Gaetano, et al. "Detect-and-segment: A deep learning approach to automate wound image segmentation." Informatics in Medicine Unlocked 29 (2022): 100884.
- Lee, Solam, et al. "Clinically applicable deep learning framework for measurement of the extent of hair loss in patients with alopecia areata." JAMA dermatology 156.9 (2020): 1018-1020.
- Gudobba, Cameron, et al. "Automating hair loss labels for universally scoring alopecia from images: rethinking alopecia scores." JAMA dermatology 159.2 (2023): 143-150.
- Xu, Rongtao, et al. "Skinformer: Learning statistical texture representation with transformer for skin lesion segmentation." IEEE Journal of Biomedical and Health Informatics 28.10 (2024): 6008-6018.
- L.-C. Chen, Y. Zhu, G. Papandreou, F. Schroff, and H. Adam, “Encoder- decoder with atrous separable convolution for semantic image segmen- tation,” in Proceedings of the European conference on computer vision (ECCV), 2018, pp. 801–818.
- Z. Zhou, M. M. Rahman Siddiquee, N. Tajbakhsh, and J. Liang, “Unet++: A nested u-net architecture for medical image segmentation,” in Deep learning in medical image analysis and multimodal learning for clinical decision support. Springer, 2018, pp. 3–11.
- Z. Gu, J. Cheng, H. Fu, K. Zhou, H. Hao, Y. Zhao, T. Zhang, S. Gao, and J. Liu, “Ce-net: Context encoder network for 2d medical image segmentation,” IEEE Transactions on Medical Imaging, vol. 38, no. 10, pp. 2281–2292, 2019.
- J. M. J. Valanarasu, P. Oza, I. Hacihaliloglu, and V. M. Patel, “Medical transformer: Gated axial-attention for medical image segmentation,” in International Conference on Medical Image Computing and Computer- Assisted Intervention. Springer, 2021, pp. 36–46.
- Y. Ji, R. Zhang, H. Wang, Z. Li, L. Wu, S. Zhang, and P. Luo, “Multi- compound transformer for accurate biomedical image segmentation,” in International Conference on Medical Image Computing and Computer- Assisted Intervention. Springer, 2021, pp. 326–336.
- R. Xu, C. Wang, S. Xu, W. Meng, and X. Zhang, “Dc-net: Dual context network for 2d medical image segmentation,” in International Confer- ence on Medical Image Computing and Computer-Assisted Intervention. Springer, 2021, pp. 503–513.
- J. Wang, L. Wei, L. Wang, Q. Zhou, L. Zhu, and J. Qin, “Boundary- aware transformers for skin lesion segmentation,” in International Con- ference on Medical Image Computing and Computer-Assisted Interven- tion. Springer, 2021, pp. 206–216.
- E. Tartaglione, A. Bragagnolo, A. Fiandrotti, and M. Grangetto, “Loss- based sensitivity regularization: towards deep sparse neural networks,” Neural Networks, vol. 146, pp. 230–237, 2022.
- D. Dai, C. Dong, S. Xu, Q. Yan, Z. Li, C. Zhang, and N. Luo, “Ms red: A novel multi-scale residual encoding and decoding network for skin lesion segmentation,” Medical Image Analysis, vol. 75, p. 102293, 2022.
- E. K. Aghdam, R. Azad, M. Zarvani, and D. Merhof, “Attention swin u- net: Cross-contextual attention mechanism for skin lesion segmentation,” in 2023 IEEE 20th International Symposium on Biomedical Imaging (ISBI). IEEE, 2023, pp. 1–5.
- Hashemifard K, Florez-Revuelta F. From garment to skin: the visuaal skin segmentation dataset. InInternational conference on image analysis and processing 2022 May 23 (pp. 59-70). Cham: Springer International Publishing.
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional network. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017
- Abubaker M, Alsadder Z, Abdelhaq H, Boltes M, Alia A. RPEE-Heads Benchmark: A Dataset and Empirical Comparison of Deep Learning Algorithms for Pedestrian Head Detection in Crowds. IEEE Access. 2025 Apr 22.
- Al-Hamadani S. Intelligent Healthcare Imaging Platform: A VLM-Based Framework for Automated Medical Image Analysis and Clinical Report Generation. arXiv preprint arXiv:2509.13590. 2025 Sep 16.
- Fagertun J, Harder S, Rosengren A, Moeller C, Werge T, Paulsen RR, Hansen TF. 3D facial landmarks: Inter-operator variability of manual annotation. BMC medical imaging. 2014 Oct 11;14(1):35.
- Hammadi Y, Grondin F, Ferland F, Lebel K. Evaluation of various state of the art head pose estimation algorithms for clinical scenarios. Sensors. 2022 Sep 10;22(18):6850.
- Huynh QT, Nguyen PH, Le HX, Ngo LT, Trinh NT, Tran MT, Nguyen HT, Vu NT, Nguyen AT, Suda K, Tsuji K. Automatic acne object detection and acne severity grading using smartphone images and artificial intelligence. Diagnostics. 2022 Aug 3;12(8):1879.
- Mac Carthy T, Montilla IH, Aguilar A, Castro RG, Pérez AM, Sueiro AV, de la Campa LV, Alfageme F, Medela A. Automatic Urticaria Activity Score: Deep Learning–Based Automatic Hive Counting for Urticaria Severity Assessment. JID Innovations. 2024 Jan 1;4(1):100218.
- Min S, Kong HJ, Yoon C, Kim HC, Suh DH. Development and evaluation of an automatic acne lesion detection program using digital image processing. Skin Research and Technology. 2013 Feb;19(1):e423-32.
- Ovadja ZN, Schuit MM, van der Horst CM, Lapid O. Inter‐and intrarater reliability of Hurley staging for hidradenitis suppurativa. British Journal of Dermatology. 2019 Aug 1;181(2):344-9.
- Park, H.; Siosund, L.; Yoo, Y.; Monet, N.; Bang, J.; Kwak, N. Sinet: Extreme lightweight portrait segmentation networks with spatial squeeze module and information blocking decoder. In Proceedings of the IEEE/CVF Winter Conference on Applications of Computer Vision, Snowmass Village, CO, USA, 1–5 March 2020
- Rashataprucksa, K., Chuangchaichatchavarn, C., Triukose, S., Nitinawarat, S., Pongprutthipan, M. and Piromsopa, K., 2020, August. Acne detection with deep neural networks. In Proceedings of the 2020 2nd International Conference on Image Processing and Machine Vision (pp. 53-56).
- Rikhye RV, Hong GE, Singh P, Smith MA, Loh A, Muralidharan V, Wong D, Sayres R, Phung M, Betancourt N, Fong B. Differences between patient and clinician-taken images: implications for virtual care of skin conditions. Mayo Clinic Proceedings: Digital Health. 2024 Mar 1;2(1):107-18.
- Sangha A, Rizvi M. Detection of acne by deep learning object detection. medRxiv. 2021 Dec 11:2021-12.
- Schultheis M, Staubach‐Renz P, Grabbe S, Hennig K, Khoury F, Nikolakis G, Kirschner U. Can hidradenitis suppurativa patients classify their lesions by means of a digital lesion identification scheme?. JDDG: Journal der Deutschen Dermatologischen Gesellschaft. 2023 Jan;21(1):27-32.
- Sui Y, Shan X, Dai L, Jing H, Li B, Ma J. S 2 Head: Small-Size Human Head Detection Algorithm By Improved YOLOv8n Architecture. IEEE Access. 2025 Aug 7.
- Takwale A, Arthur E, Pearce J, Farrant P, Holmes S, Harries M. A practical guide to the standardization of hair loss photography for clinicians. Clinical and Experimental Dermatology. 2025 Mar;50(3):564-72.
- Thorlacius L, Garg A, Riis PT, Nielsen SM, Bettoli V, Ingram JR, Del Marmol V, Matusiak L, Pascual JC, Revuz J, Sartorius K. Inter‐rater agreement and reliability of outcome measurement instruments and staging systems used in hidradenitis suppurativa. British Journal of Dermatology. 2019 Sep 1;181(3):483-91.
- Wen H, Yu W, Wu Y, Zhao J, Liu X, Kuang Z, Fan R. Acne detection and severity evaluation with interpretable convolutional neural network models. Technology and Health Care. 2022 Jan;30(1_suppl):143-53.
- Xu H, Sarkar A, Abbott AL. Color invariant skin segmentation. InProceedings of the IEEE/CVF conference on computer vision and pattern recognition 2022 (pp. 2906-2915).
- You H, Lee K, Oh J, Lee EC. Efficient and low color information dependency skin segmentation model. Mathematics. 2023 Apr 26;11(9):2057.
- Zhang Z, Xia S, Cai Y, Yang C, Zeng S. A soft-YoloV4 for high-performance head detection and counting. Mathematics. 2021 Nov 30;9(23):3096.
Traceability to QMS Records
Signature meaning
The signatures for the approval process of this document can be found in the verified commits at the repository for the QMS. As a reference, the team members who are expected to participate in this document and their roles in the approval process, as defined in Annex I Responsibility Matrix of the GP-001, are:
- Author: Team members involved
- Reviewer: JD-003 Design & Development Manager, JD-004 Quality Manager & Person Responsible for Regulatory Compliance (PRRC)
- Approver: JD-001 General Manager