R-TF-028-005 AI Development Report
Table of contents
- Introduction
- Data Management
- Model Development and Validation
- Summary and Conclusion
- State of the Art Compliance and Development Lifecycle
- Integration Verification Package
- Purpose
- Package Location and Structure
- Package Contents Per Model
- Acceptance Criteria
- Model-Specific Package Details
- Clinical Models - ICD Classification and Binary Indicators
- Clinical Models - Visual Sign Intensity Quantification
- Clinical Models - Wound Characteristic Assessment
- Clinical Models - Lesion Quantification
- Clinical Models - Surface Area Quantification
- Clinical Models - Pattern Identification
- Non-Clinical Models
- Verification Procedure for Software Integration Team
- Traceability
- AI Risks Assessment Report
- Related Documents
Introduction
Context
This report documents the development, verification, and validation of the AI algorithm package for the Legit.Health Plus medical device. The development process was conducted in accordance with the procedures outlined in GP-028 AI Development and followed the methodologies specified in the R-TF-028-002 AI Development Plan.
The algorithms are designed as offline (static) models. They were trained on a fixed dataset prior to release and do not adapt or learn from new data after deployment. This ensures predictable and consistent performance in the intended environment for non-clinical attribute analysis.
Algorithms Description
The Legit.Health Plus device incorporates a suite of AI models that support the device's intended purpose, including a set of non-clinical models focused on the analysis of non-clinical image attributes. These non-clinical models are essential for ensuring the quality, suitability, and proper categorization of dermatological images, and for supporting downstream clinical and research workflows. A comprehensive description of all models, their objectives, and performance specifications is provided in R-TF-028-001 AI/ML Description.
Non-Clinical Models (supporting proper functioning and non-clinical data analysis):
- Acneiform Inflammatory Pattern Identification: Translates objective lesion counts and density into standardized IGA severity scores, supporting consistent acne severity assessment.
- Skin Tone Identification: Assigns Fitzpatrick and Monk skin tone labels to images for demographic analysis and bias monitoring.
- Body Site Identification: Classifies anatomical locations visible in the image to support site-specific analysis and data organization.
- Skin 3D Reconstruction: Transforms 2D image data into 3D spatial coordinates for advanced surface and volume quantification.
- Color Correction: Standardizes color representation in images using reference markers, ensuring consistency across varying lighting and camera conditions.
This report focuses on the development methodology, data management processes, and validation results for all non-clinical models. Each model shares a common data foundation but may require specific annotation procedures as detailed in the respective data annotation instructions.
AI Standalone Evaluation Objectives
The standalone validation aimed to confirm that all non-clinical AI models meet their predefined performance criteria as outlined in R-TF-028-001 AI/ML Description.
Performance specifications and success criteria vary by model type and are detailed in the individual model sections of this report. All non-clinical models were evaluated on independent, held-out test sets that were not used during training or model selection.
Data Management
Overview
The development of all AI models in the Legit.Health Plus device relies on a comprehensive dataset compiled from multiple sources and annotated through a multi-stage process. This section describes the general data management workflow that applies to all models, including collection, foundational annotation (ICD-11 mapping), and partitioning. Model-specific annotation procedures are detailed in the individual model sections.
Data Collection
The dataset was compiled from multiple distinct sources:
- Archive Data: Images sourced from reputable online sources and private institutions, as detailed in
R-TF-028-003 Data Collection Instructions - Archive Data. - Custom Gathered Data: Images collected under formal protocols at clinical sites, as detailed in
R-TF-028-003 Data Collection Instructions - Custom Gathered Data.
This combined approach resulted in a comprehensive dataset covering diverse demographic characteristics (age, sex, Fitzpatrick skin types I-VI), anatomical sites, imaging conditions, and pathological conditions.
Dataset summary:
| Item | Value |
|---|---|
| Total ICD-11 categories | 850 |
| Total images | 280342 |
| Images of FST-1 | 89225 (31.83%) |
| Images of FST-2 | 91349 (32.58%) |
| Images of FST-3 | 59610 (21.26%) |
| Images of FST-4 | 23466 (8.37%) |
| Images of FST-5 | 11914 (4.25%) |
| Images of FST-6 | 4778 (1.70%) |
| Images of female | 52857 (18.85%) |
| Images of male | 55334 (19.74%) |
| Images of unspecified sex | 172151 (61.41%) |
| Images of Pediatric | 12829 (4.58%) |
| Images of Adult | 52694 (18.80%) |
| Images of Geriatric | 28350 (10.11%) |
| Images of unspecified age | 186469 (66.51%) |
| ID | Dataset Name | Type | Description | ICD-11 Mapping | Crops | Diff. Dx | Sex | Age |
|---|---|---|---|---|---|---|---|---|
| 1 | Torrejon-HCP-diverse-conditions | Multiple | Dataset of skin images by physicians with good photographic skills | ✓ Yes | Varies | ✓ | ✓ | ✓ |
| 2 | Abdominal-skin | Archive | Small dataset of abdominal pictures with segmentation masks for `Non-specific lesion` class | ✗ No | Yes (programmatic) | — | — | — |
| 3 | Basurto-Cruces-Melanoma | Custom gathered | Clinical validation study dataset (`MC EVCDAO 2019`) | ✓ Yes | Yes (in-house crops) | — | ✓ | ✓ |
| 4 | BI-GPP (batch 1) | Archive | Small set of GPP images from Boehringer Ingelheim (first batch) | ✓ Yes | No | — | — | — |
| 5 | BI-GPP (batch 2) | Archive | Large dataset of GPP images from Boehringer Ingelheim (second batch) | ✓ Yes | Yes (programmatic) | — | ✓ | ✓ |
| 6 | Chiesa-dataset | Archive | Sample of head and neck lesions (Medela et al., 2024) | ✓ Yes | Yes (in-house crops) | — | ◐ | ◐ |
| 7 | Figaro 1K | Archive | Hair style classification and segmentation dataset, repurposed for `Non-specific finding` | ✗ No | Yes (in-house crops) | — | — | — |
| 8 | Hand Gesture Recognition (HGR) | Archive | Small dataset of hands repurposed for non-specific images | ✗ No | Yes (programmatic) | — | — | — |
| 9 | IDEI 2024 (pigmented) | Archive | Prospective and retrospective studies at IDEI (DERMATIA project), pigmented lesions only | ✓ Yes | Yes (programmatic) | — | ✓ | ◐ |
| 10 | Manises-HS | Archive | Large collection of hidradenitis suppurativa images | ✗ No | Not yet | — | ✓ | ✓ |
| 11 | Nails segmentation | Archive | Small nail segmentation dataset repurposed for `non-specific lesion` | ✗ No | Yes (programmatic) | — | — | — |
| 12 | Non-specific lesion V2 | Archive | Small representative collection repurposed for `non-specific lesion` | ✗ No | Yes (programmatic) | — | — | — |
| 13 | Osakidetza-derivation | Archive | Clinical validation study dataset (`DAO Derivación O 2022`) | ✓ Yes | Yes (in-house crops) | ◐ | ✓ | ✓ |
| 14 | Ribera ulcers | Archive | Collection of ulcer images from Ribera Salud | ✗ No | Yes (from wound masks, not all) | — | — | — |
| 15 | Transient Biometrics Nails V1 | Archive | Biometric dataset of nail images | ✗ No | Yes (programmatic) | — | — | — |
| 16 | Transient Biometrics Nails V2 | Archive | Biometric dataset of nail images | ✗ No | No (close-ups) | — | — | — |
| 17 | WoundsDB | Archive | Small chronic wounds database | ✓ Yes | No | — | ✓ | ◐ |
| 18 | Clinica Dermatologica Internacional - Acne | Custom gathered | Compilation of images from CDI's acne patients with IGA labels | ✓ Yes | No | — | — | — |
| 19 | Manises-DX | Archive | Large collection of images of different dermatological categories | ✓ Yes | Not yet | — | — | — |
Total datasets: 55 | With ICD-11 mapping: 41
Legend: ✓ = Yes | ◐ = Partial/Pending | — = No
Model Development and Validation
This section details the development, training, and validation of all non-clinical AI models in the Legit.Health Plus device. Each model subsection includes:
- Model-specific data annotation requirements
- Training methodology and architecture
- Performance evaluation results
- Bias analysis and fairness considerations
Acneiform Inflammatory Pattern Identification
Model Overview
Reference: R-TF-028-001 AI/ML Description - Acneiform Inflammatory Pattern Identification section
This Acneiform Inflammatory Pattern Identification model assess the intensity of the acneiform inflammatory lesion affection by analyzing tabular features derived from the Acneiform Inflammatory Lesion Quantification algorithm. This model is a mathematical equation designed and optimized with state-of-the-art evolutionary and optimization algorithms to maximize its correlation with the Investigator's Global Assessment (IGA) scores consensued from expert dermatologists criteria.
Clinical Significance: severity assessment aids in achieving a more objective evaluation and select the most appropriate treatment plan.
Data Requirements and Annotation
Foundational annotation: ICD-11 mapping. Images belonging to the Clinica Dermatologica Internacional - Acne subset.
Model-specific annotation: Intensity annotation (R-TF-028-004 Data Annotation Instructions)
Images are annotated by a panel of three dermatologists with expertise in acneiform pathologies. Annotations consist of IGA scores ranging from 0 (clear skin) to 4 (severe acne). The final label for each image is determined by the mathematical consensus among the three dermatologists.
Dataset statistics:
- Images with acneiform lesions: 331, including diverse types of acneiform inflammatory lesions (e.g., papules, pustules, comedones) obtained from the main dataset.
- All images picture faces profiles and not specific close-ups of the lesions.
- Number of subjects: ~165 (estimated)*
- Average inter-annotator Pearson coefficient variability: 0.56 (0.39-0.68)
- Since the model is a mathematical equation optimized with evolutionary algorithms rather than a deep neural network, it does not require a conventional train/validation split. This approach is considerably less data-dependent than deep learning models, making the 331-image dataset sufficient for robustly tune and validate the limited number of parameters in the formula.
*Subject count estimation methodology: Due to the nature of this single-source clinical dataset (Clinica Dermatologica Internacional - Acne), explicit subject-level identifiers were not preserved in the anonymized dataset. The estimated subject count was derived through systematic visual review of facial profile images to identify potential duplicate subjects based on anatomical features and lesion patterns. A conservative estimation factor of 2.0 images per subject was applied, consistent with clinical photography protocols that typically capture multiple angles per patient visit. This estimation was validated through independent review by two data analysts and is subject to a margin of error of approximately ±20%. The optimization process uses cross-validation techniques that inherently account for potential subject overlap.
Training Methodology
Pre-processing
The Acneiform Inflammatory Pattern Identification model analyzes the detection data derived from the Acneiform Inflammatory Lesion Quantification model. This detection data is pre-processed to generate features that include the counts of acneiform inflammatory lesions detected in the image () and their density ().
Given the localization and dimensions of the detected acneiform lesions, we define the density as the ratio between the overlapping detection area and the total area covered by the detected lesions. Prior to computing density, we pre-process detections by first converting the detected rectangular bounding boxes into circles and enlarging their radius by a factor of 6. The circular transformation refines lesion boundaries by excluding irrelevant bounding box corners, while enlargement improves collision detection among closely positioned lesions. This density score is 0-1 bounded.
Model design and optimization
The Acneiform Inflammatory Pattern Identification formula is designed using a two-stage semi-automatic process based on symbolic learning and differential evolution optimization. The symbolic learning stage generates a set of mathematical expressions that combine both and with different mathematical operands. In the second stage, we add constant variables (a and b weights) when possible to the mathematical expressions and refine them with differential evolution optimization. This entire optimization process is designed to maximize the Cohen’s Kappa score by aligning the Acneiform Inflammatory Pattern Identification model's output with the IGA scores consensued from the expert dermatologists's criteria. As a result of this process, we set the Acneiform Inflammatory Pattern Identification model as the formula with the best trade-off between performance and interpretability. The model's scores grow with the number of lesions and density.
Final Acneiform Inflammatory Pattern Identification model is with parameters and .
For the symbolic learning stage, we use the PySR Python library with 250 iterations, 50 populations, population size 50, maxsize 10, and L2 loss. For the differential evolution optimization stage, we use the SciPy Python library for 10000 iterations (remaining hyperparameters set as default).
Post-processing
Equation scores are clipped to a maximum of 4 to align with the IGA scale (0-4), and later weighted x2.5 to map the output to a 0-10 scale, providing a more granular severity assessment.
Performance Results
Performance is evaluated using the Pearson correlation coefficient, to account for the similarity between model outputs and expert IGA scores. Statistics are calculated with 95% confidence intervals using bootstrapping (1000 samples). Success criteria is defined as Pearson ≥ 0.39 to account for a performance non-inferior to the expert inter-annotator variability.
| Metric | Result | Success Criterion | Outcome |
|---|---|---|---|
| Pearson | 0.55 (0.46-0.63) | ≥ 0.39 | PASS |
Verification and Validation Protocol
Test Design:
- Annotations consensued from medical expert criteria are used as gold standard for validation.
- Evaluation across diverse skin tones.
- The set of evaluation images has been extended with 40 new images. These images are sourced from the main dataset and were translated semi-automatically to darker Fitzpatrick skin types (V-VI) with the Nano Banana AI-tool. These images preserve the acneiform inflammatory lesions but with a darker skin tone.
Complete Test Protocol:
- Input: acneiform inflammatory lesion detections calculated for the validation set by the Acneiform Inflammatory Lesion Quantification model.
- Pre-processing: Feature extraction (lesion count and density).
- Processing: Score clipping and scaling.
- Output: Predicted intensity scores.
- Reference standard: Expert-annotated IGA scores.
- Statistical analysis: Pearson correlation coefficient.
Data Analysis Methods:
- Pearson correlation coefficient between model outputs and expert IGA scores.
Test Conclusions:
- The model met all success criteria, demonstrating reliable acneiform inflammatory pattern identification and suitable for clinical acne severity assessment.
- The model demonstrates non-inferiority with respect to expert annotators.
- The model's performance is within acceptable limits.
- The model showed robustness across different skin tones and severities, indicating generalizability.
Bias Analysis and Fairness Evaluation
Objective: Ensure acneiform inflammatory pattern identification performs consistently across demographic subpopulations.
Subpopulation Analysis Protocol:
1. Fitzpatrick Skin Type Analysis:
- Performance stratified by Fitzpatrick skin types: I-II (light), III-IV (medium), V-VI (dark).
- Success criterion: Pearson ≥ 0.39.
| Subpopulation | Num. validation images | Pearson | Outcome |
|---|---|---|---|
| Fitzpatrick I-II | 282 | 0.57 (0.48-0.66) | PASS |
| Fitzpatrick III-IV | 49 | 0.51 (0.27-0.69) | PASS |
| Fitzpatrick V-VI | 40 | 0.55 (0.26-0.76) | PASS |
Results Summary:
- The model demonstrated reliable mean performance across Fitzpatrick skin types I-VI, meeting all success criteria.
- Confidence intervals for the Fitzpatrick III-IV and V-VI groups are wider due to smaller sample size, slightly exceeding the inter-annotator variability range.
Bias Mitigation Strategies:
- Image augmentation including color and lighting variations during training.
- Pre-training on diverse data to improve generalization.
Bias Analysis Conclusion:
- The model demonstrated consistent performance across Fitzpatrick skin types, with all success criteria met.
- Continued efforts to collect diverse data, especially for underrepresented groups, will further enhance model robustness and fairness and provide more robust evaluation statistics.
Skin Tone Identification
Model Overview
Reference: R-TF-028-001 AI/ML Description - Skin Tone Identification
This model classifies skin into Fitzpatrick (I-VI) and Monk (0-9) tones to enable bias monitoring and fairness evaluation across all clinical models.
Clinical Significance: Essential for ensuring equitable performance across diverse patient populations and detecting potential algorithmic bias.
Data Requirements and Annotation
Compiled dataset: 36563 clinical and non-clinical images sourced from:
representative-expert-annotated-set, composed by clinical and non-clinical images from the subsets11kHands,smartskins,blackandbrownskin,acne04,SALT-batch3,PAD-UFES-20,Abdominal skin segmentation, andDDIFitzpatrick17kHumanaeMST-ESCIN
Model-specific annotation: Skin Tone Identification (R-TF-028-024 Data Annotation Instructions)
Annotatios are sourced from each dataset's original annotations if available and valid, or annotated by experts. The annotations are:
- Fitzpatrick skin tones: One of six categories, from I to VI.
- Monk skin tones: One of ten categories, from 0 to 9.
Dataset statistics:
The dataset is split in train, validation, and test sets. The dataset is split at patient level to avoid data leakage when subject information is available.
- Images: 36563
- Images can be clinical or non-clinical and span a broad range of skin conditions, body parts, and skin tones.
- 35074 images with valid fitzpatrick annotations. 36563 images with valid monk annotations.
- Train, validation, and test set contain 27343, 4805, and 4415 images respectively.
- Images annotated by experts are annotated by 3 persons. Final labels are the mathematical consensus of the 3 annotations.
- The inter-rater variability of the
representative-expert-annotated-sethas Fitzpatrick Accuracy of 0.41 (0.34-0.49) and MAE of 0.70 (0.57-0.81). - The inter-rater variability of the
representative-expert-annotated-sethas Monk Accuracy of 0.42 (0.37-0.47) and MAE of 0.74 (0.67-0.82).
Training Methodology
The model architecture and training hyperparameters were selected after a systematic hyperparameter tuning process. We compared different image encoders (e.g., ConvNext and EfficientNet of different sizes) and evaluated multiple data hyperparameters (e.g., input resolutions, augmentation strategies) and optimization configurations (e.g., batch size, learning rate, metric learning). The final configuration was chosen as the best trade-off between performance and runtime efficiency.
Architecture:
The model is a multi-task neural network designed to predict Fitzpatrick and Monk skin tone scales simultaneously, while also generating embeddings for metric learning. It uses a shared backbone and common projection head, branching into specific heads for each skin tone scale.
- Backbone (Encoder):
- Model: ConvNext Small, pre-trained on the ImageNet dataset.
- Regularization: dropout and drop path.
- Common Projection Head:
- A shared processing block that maps encoder features to a common latent space of 256 features.
- Consists of a ReLU activation, Dropout, and a Linear layer.
- Task-Specific Heads: The model splits into two distinct branches, one for Fitzpatrick and one for Monk skin tone scales. Each branch receives the 256-dimensional output from the Common Projection Head and contains two sub-heads:
- Classification Head:
- A dedicated block (ReLU, Dropout, Linear)
- Output size: 6 classes for Fitzpatrick (I-VI) and 10 classes for Monk (0-9).
- Metric Embedding Head:
- A multi-layer perceptron (two sequential blocks of ReLU, Dropout, and Linear layers) that outputs feature embeddings.
- Output size: 256 features.
- Classification Head:
- Weight Initialization:
- Linear Layers: Kaiming Normal initialization.
- Biases: Initialized to zero.
The model is implemented with PyTorch and the Python timm library.
Training approach:
The training process employs a multi-task learning strategy with Dynamic Weight Averaging (DWA), optimizing for both classification accuracy and embedding quality across Fitzpatrick and Monk skin tone scales. It utilizes a two-stage approach, starting with a frozen backbone followed by full model fine-tuning. It also incorporates data augmentation and mixed-precision training.
- Training Stages:
- Stage 1 (Frozen Backbone): Trains only the projection and task-specific heads for 7 epochs.
- Stage 2 (Fine-tuning): Trains the entire model for 25 epochs.
- Optimization:
- Optimizer: AdamW Schedule-Free with weight decay (0.01).
- Base LR: 0.0025
- Learning Rate: Includes a 4-epoch warmup. During fine-tuning (Stage 2), the base learning rate is scaled by 0.4x, and the backbone learning rate is further scaled down (0.4x) relative to the task-specific heads.
- Gradient Clipping: Gradients are clipped to a norm of 1.0.
- Precision: Mixed precision training using BFloat16.
- Loss Functions:
- Classification: Cross-Entropy Loss with inverse class frequency weighting to handle class imbalance.
- Metric Learning: NTXentLoss for embedding quality.
- Loss Weighting: Dynamic Weight Averaging (DWA) with temperature T=1.0 and scaling enabled, balancing the four losses (Fitzpatrick classification, Fitzpatrick metric, Monk classification, Monk metric).
Pre-processing:
- Augmentation: Geometric transformations, contrast and saturation transformations, and coarse dropout.
- Input: Images are resized to 384x384 pixels with a batch size of 64.
Post-processing:
- Classification probabilities are computed applying the softmax operation over the classification logits.
- Classification categories are selected as the ones with higher probability.
Performance Results
Performance is evaluated using accuracy and Mean Absolute Error (MAE) to account for the correct Fitzpatrick and Monk skin tone scales. Succeed critera is set as accuracy ≥ 41% and MAE ≤ 1 for Fitzpatrick and accuracy ≥ 42% and MAE ≤ 1 for Monk. Succed criteria accounts for a model performance on par to the inter-rater performance of expert annotators.
| Metric | Result | Success Criterion | Outcome |
|---|---|---|---|
| Fitzpatrick Accuracy | 0.62 (0.58-0.65) | ≥ 0.41 | PASS |
| Fitzpatrick MAE | 0.39 (0.35-0.44) | ≤ 1 | PASS |
| Monk Accuracy | 0.54 (0.50-0.58) | ≥ 0.42 | PASS |
| Monk MAE | 0.49 (0.45-0.53) | ≤ 1 | PASS |
Verification and Validation Protocol
Test Design:
- 640 images from the test split of the
representative-expert-annotated-setdataset. - Images are annotated with 3 expert annotators. Annotations consensued from expert criteria are used as gold standard for validation.
- Clinical and non-clinical images representing diverse anatomical sites, lighting conditions, and skin tone spectrum.
Complete Test Protocol:
- Input: Images of skin from the test set.
- Pre-processing: Image resizing to 384x384 pixels and normalization.
- Processing: Skin tone classification model inference.
- Output: Predicted Fitzpatrick tone (I-VI) and Monk skin tone (0-9) with confidence scores.
- Ground truth: Expert-annotated skin tone scales (mathematical consensus from 3 annotators).
- Statistical analysis: Accuracy, Mean Absolute Error (MAE).
Data Analysis Methods:
- Classification accuracy.
- Mean Absolute Error (MAE).
- Confusion matrix.
Test Conclusions:
- The model met all success criteria, demonstrating reliable skin tone identification suitable for bias monitoring and fairness evaluation.
- The model demonstrates non-inferiority with respect to expert annotators.
- The model's performance is within acceptable limits.
- The model showed robustness across different datasets and imaging conditions, indicating generalizability.
Bias Analysis and Fairness Evaluation
Objective: This model itself is a bias mitigation tool. Validation ensures accurate identification across the full Fitzpatrick spectrum.
Subpopulation Analysis Protocol:
1. Fitzpatrick Skin Tone Analysis:
- Performance stratified by Fitzpatrick skin tones: I-II (light), III-IV (medium), V-VI (dark).
- Metrics evaluated: Accuracy and Mean Absolute Error (MAE).
- Fitzpatrick success criteria: Accuracy ≥ 0.41; MAE ≤ 1.
- Monk success criteria: Accuracy ≥ 0.42; MAE ≤ 1.
| Subpopulation | Num. train images | Num. val images | Num. test images | Fitzpatrick Acc | Fitzpatrick MAE | Monk Acc | Monk MAE | Outcome |
|---|---|---|---|---|---|---|---|---|
| Fitzpatrick I-II | 14627 | 2391 | 382 | 0.55 (0.50-0.59) | 0.47 (0.42-0.52) | 0.54 (0.48-0.61) | 0.49 (0.42-0.56) | PASS |
| Fitzpatrick III-IV | 8886 | 1482 | 130 | 0.65 (0.57-0.72) | 0.35 (0.28-0.43) | 0.53 (0.47-0.59) | 0.50 (0.43-0.57) | PASS |
| Fitzpatrick V-VI | 2799 | 474 | 14 | 0.86 (0.64-1.00) | 0.14 (0.00-0.36) | 0.56 (0.45-0.67) | 0.45 (0.33-0.58) | PASS |
Results Summary:
- The model met all success criteria, demonstrating reliable skin tone identification suitable for bias monitoring and fairness evaluation.
- The model presents consistent robustness across all skin tone subpopulations, for both Fitzpatrick and Monk scales.
- The model demonstrates non-inferiority with respect to expert annotators.
- The model's performance is within acceptable limits.
Bias Mitigation Strategies:
- Image augmentation including gemetric, contrast, saturation and dropout augmentations.
- Class-balancing to ensure equal representation of all classes.
- Use of metric learning to improve the model's ability to generalize to new data.
- Pre-training on diverse data to improve generalization
- Two-stage training to fit the model to the new data while benefiting from the image encoder pre-training.
Bias Analysis Conclusion:
- The model demonstrated consistent performance across Fitzpatrick skin tones, for both Fitzpatrick and Monk scales.
- The model met all success criteria, demonstrating reliable skin tone identification suitable for bias monitoring and fairness evaluation.
Skin 3D Reconstruction
Model Overview
Reference: R-TF-028-001 AI/ML Description - Skin 3D Reconstruction
This framework transforms 2D pixel coordinates from a standard 2D image into 3D world metric coordinates, enabling comprehensive and accurate spatial analysis of skin surfaces. This methodology allows for straightforward geometric analysis, including the calculation of the area, perimeter, axes, volume, and depth of target surfaces.
Clinical Significance: Accurate surface area quantification is fundamental to severity scoring in dermatology, including calculations for PASI, EASI, VASI, and BSA. It allows clinicians to monitor lesion sizes over time, assess condition severity, and determine precise treatment dosages. By accounting for depth variation, this method significantly reduces measurement errors associated with perspective distortion, camera angles, and irregular body surface curvatures.
Data Requirements and Annotation
Model-specific annotation: Extent Annotation (R-TF-028-004 Data Annotation Instructions).
To evaluate the method's precision, we designed a controlled experimental setup to simulate real-world clinical use cases with a precise gold standard. We drew squares of known dimensions (15 mm per side; 225 mm² area) directly onto the skin to serve as the ground truth. The images captured these target surfaces alongside 5 reference markers (10 mm side) arranged in a 5x5 grid pattern.
Dataset statistics:
- Images evaluated: 58 images captured using a commercially available smartphone (Apple iPhone 15 Pro Max).
- Reference shapes: 410 reference shapes of known dimensions.
- Subjects: 10 different white volunteers.
- Anatomical locations: Face, neck, and foot.
- Camera configurations: Focal lengths of 24mm, 35mm, and their ultra-wide variations.
- Real-world artifacts present: High and shallow hair density, makeup, jewelry, and freckles.
Methodology
The solution quantifies the surface of a skin region delineated with a list of pixels. It leverages state-of-the-art monocular relative depth estimation, world reference information, and camera calibration parameters. During the development phase, we rigorously tested several variations of this framework to identify the best configuration. This included testing alternative depth estimation methods and assessing markers of different sizes, shapes, and quantities (from 2 to 5 markers) to maximize accuracy while minimizing clinical hassle.
Architecture and framework pipeline:
- Relative Depth Estimation: The framework utilizes
Depth Anything v2(ViT-B version), a state-of-the-art monocular relative depth estimation model, trained on 42 million images to generate highly detailed relative depth maps. - Metric Depth Scaling: Because the depth model outputs relative values, we apply a linear transformation to convert these values to metric measurements (millimeters). We optimize the scale and translation parameters of this transformation using the Levenberg-Marquardt algorithm and the reference information of the physical markers placed around the target body surface.
- 3D Coordinate Transformation: For any given pixel coordinate , we compute the 3D metric world coordinates using the camera's intrinsic parameters and the calculated metric depth :
Surface quantification endpoints:
- Area calculation: We break down the target surface into a set of smaller, non-overlapping planar triangles using the Delaunay triangulation algorithm. After transforming the vertices of these triangles into 3D world coordinates, we compute the individual planar triangle areas. We calculate the total surface area by summing all individual triangular subparts.
- Perimeter calculation: We calculate the perimeter of the target surface by summing the lengths of the edges of the triangulated surface. We compute each edge length using the 3D coordinates of its vertices, providing an accurate measurement that accounts for surface curvature.
- Axes calculation: We set the major axis as the pair of points within the target surface that are furthest apart. We calculate the minor axis as the longest line segment that we can draw between two points on the surface that is (near) perpendicular to the major axis. We calculate both axes in 3D space, providing accurate measurements regardless of surface curvature or orientation.
- Depth calculation: We calculate the depth of the target surface as the maximum distance between the centroid of each axis and the 3D coordinates of the surface points. This method provides a robust depth estimation that accounts for surface curvature and orientation, as it considers the spatial distribution of all points on the surface rather than relying on a single point.
- Volume calculation: We calculate the volume of the target surface by summing the volumes of the tetrahedrons formed by the 3D coordinates of the triangulated surface when we project them onto the reference plane defined by 3 points extracted from the axes limits. This method provides an accurate volume estimation that accounts for surface curvature and orientation, as it considers the spatial distribution of all points on the surface rather than relying on a single point.
Performance Results
Performance is evaluated using the Relative Mean Absolute Error (rMAE) compared against the expected ground truth areas (225 mm²). Success criterion: rMAE to account for a strict tolerance.
| Metric | Result | Success Criterion | Outcome |
|---|---|---|---|
| rMAE | 0.09 (0.08-0.10) | PASS |
Verification and Validation Protocol
Test Design:
- Evaluation against a precisely defined gold standard utilizing geometric shapes of 225 mm² drawn on the skin.
- Testing conducted across heterogeneous body parts with high anatomical complexity (face, neck, foot).
- Robustness assessed against variations in camera configurations and body site characteristics.
Complete Test Protocol:
- Input: 2D smartphone images containing the targeted skin surfaces and specific physical reference markers.
- Processing: Delineation of the target surface, extraction of the depth map, transformation to 3D metric coordinates using marker scaling, and Delaunay triangulation.
- Output: Measured total area in metric units (mm²).
- Reference standard: Expected area of 225 mm².
Data Analysis Methods:
- Statistical analysis: relative Mean Absolute Error (rMAE).
- Visualization of the reconstructed 3D surfaces.
Test Conclusions:
- The proposed framework meet all success criteria, demonstrating accurate 3D reconstruction and surface area quantification suitable for clinical applications.
- The framework performance is within acceptable limits.
- The method's performance is robust across different anatomical sites, camera configurations, and real-world artifacts, indicating its generalizability and potential for widespread clinical use.
Bias Analysis and Fairness Evaluation
Objective: Ensure the 3D surface area reconstruction method functions accurately and consistently across different camera configurations and anatomical landscapes.
Subpopulation Analysis Protocol:
1. Body Part Analysis:
- Performance evaluated on complex anatomical region (Face, Neck, Foot).
- Success criterion: rMAE .
| Body part | Num. Images | Num. target shapes | rMAE | Outcome |
|---|---|---|---|---|
| Face | 20 | 178 | 0.10 (0.09-0.11) | PASS |
| Neck | 20 | 118 | 0.08 (0.07-0.09) | PASS |
| Foot | 18 | 114 | 0.09 (0.08-0.10) | PASS |
Results Summary:
- The method demonstrated reliable performance across different body parts, meeting all success criteria.
- The method's performance is robust to anatomical complexity and real-world artifacts, indicating its generalizability across diverse clinical scenarios.
2. Focal Length Analysis:
- Performance evaluated utilizing varied focal lengths (24mm, 35mm).
- Success criterion: rMAE .
| Focal length | Num. Images | Num. target shapes | rMAE | Outcome |
|---|---|---|---|---|
| 24mm | 29 | 205 | 0.09 (0.08-0.10) | PASS |
| 35mm | 29 | 205 | 0.09 (0.08-0.10) | PASS |
Results Summary:
- The method demonstrated reliable performance across different focal lengths, meeting all success criteria.
- The method's performance is robust to different camera configurations, indicating its generalizability across diverse clinical scenarios.
3. Focal Length Analysis:
- Performance evaluated utilizing ultra-wide and no ultra-wide configurations.
- Success criterion: rMAE .
| Ultra | Num. Images | Num. target shapes | rMAE | Outcome |
|---|---|---|---|---|
| Ultra | 34 | 220 | 0.09 (0.08-0.10) | PASS |
| No Ultra | 24 | 190 | 0.09 (0.08-0.10) | PASS |
Results Summary:
- The method demonstrated reliable performance across different ultra-wide configurations, meeting all success criteria.
- The method's performance is robust to different photographic perspectives, indicating its generalizability across diverse clinical scenarios.
4. Skin Tone Analysis:
- Although the evaluation is limited to the avialble 10 volunteers, we qualitatively observe the Depth Anything V2 performance on a set of images of diverse skin tones.
- These images are extracted from the Fitzpatrick17k dataset and cover all the Fitzpatrick skin tones and skin pathologies of variable depth complexities.
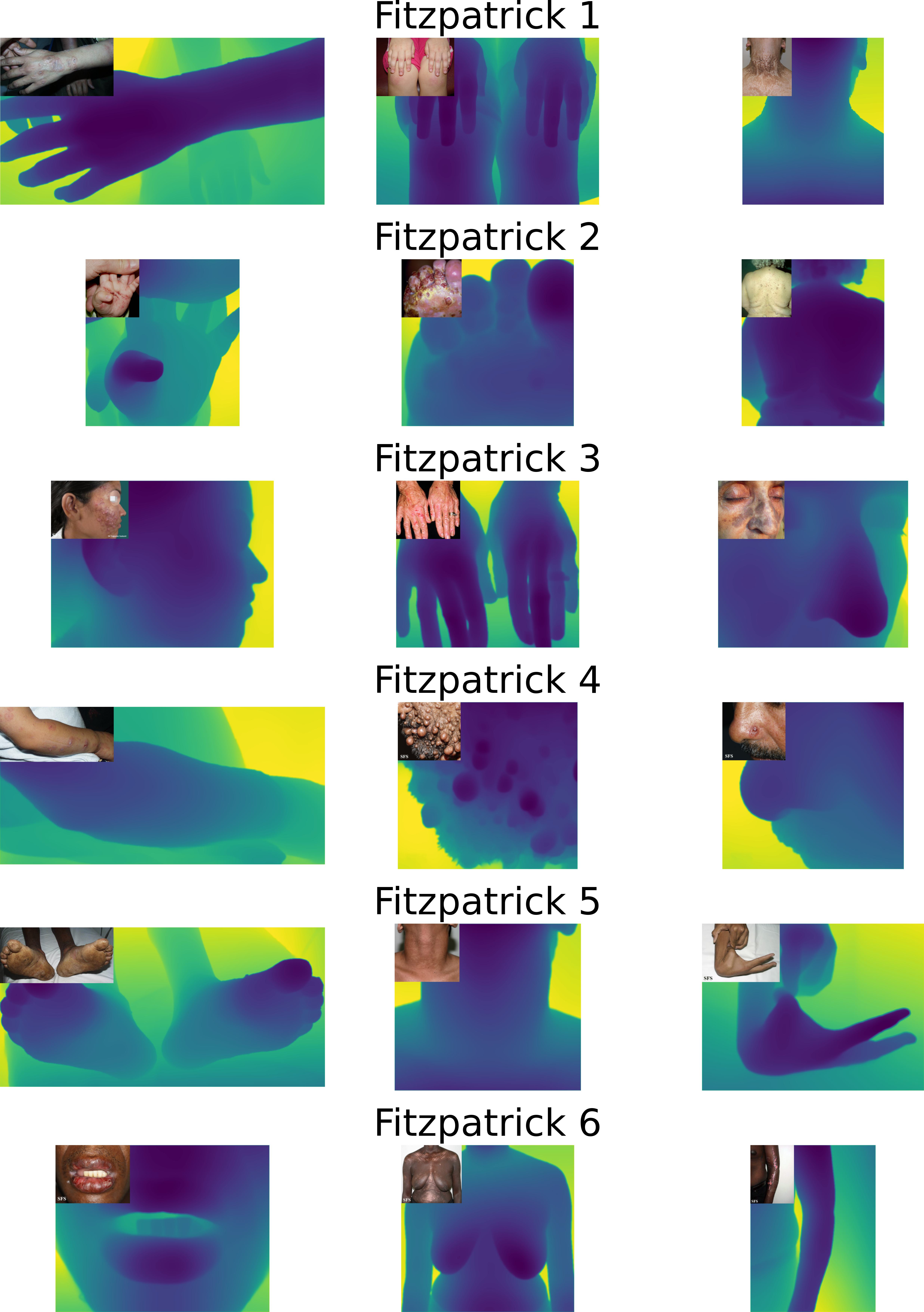 |
|---|
Relative depth map generation on a diverse set of images from the Fitzpatrick17k dataset.
- The method demonstrates consistent performance across different skin tones and pathologies, indicating its generalizability across diverse clinical scenarios.
- The use of the Depth Anything v2 model, which was trained on a highly diverse dataset, provides confidence in the method's ability to generalize across different skin tones and demographics.
Bias Mitigation Strategies:
- Utilization of a state-of-the-art Foundational Model: The core depth estimation component of the framework relies on Depth Anything v2, a robust foundational model that was trained on a massive dataset of over 42 million images. The sheer scale of this training data provides a broad representation of diverse skin tones, clinical artifacts, genders, age groups, pathologies, and varying depth complexities, which fundamentally helps mitigate inherent demographic biases. Note that we do not need to re-train this model for our specific use case, as it already demonstrates strong generalization capabilities across a wide range of scenarios, including those relevant to dermatology.
- Rigorous testing across diverse anatomical sites, camera configurations, skin tones, and pathologies to ensure consistent performance.
Bias Analysis Conclusion:
- The method demonstrated reliable performance across different body parts, camera configurations, real-world artifacts, patient demographics, and pathologies,meeting all success criteria.
- The method's performance is robust to anatomical complexity and photographic variations, indicating its generalizability across diverse clinical scenarios.
- The method shows performance within acceptable limits across all subpopulations.
Color Correction
Model Overview
Reference: R-TF-028-001 AI/ML Description - Color Correction
This preprocessing model standardizes the color representation of clinical images by detecting physical reference markers with known color values and learning a transformation to map observed colors to their theoretical ground truth.
Clinical Significance: Standardized color representation is crucial for accurate visual sign analysis, ensuring consistent performance across images captured under varying lighting conditions and camera settings. By correcting color discrepancies, this model enhances the human interpretability of images for clinical decision-making and aids in the reliability of downstream AI models that rely on color information.
Data Requirements and Annotation
Model-specific data: A curated set of images containing standardized color reference markers (5x5 grid) captured under various illuminants.
Dataset statistics:
- Total Evaluation Images: 15 captures under diverse lighting.
- Marker Types: Custom dermatological markers (5x5).
- Marker Size: 10 mm per side plus a thin white border.
- Lighting Conditions: Warm, Neutral, Cool.
Methodology
Marker Detection:
We detect markers by first identifying candidate contours using adaptive thresholding and filtering them based on geometry and the presence of a black border. Then, we extract the color values from the marker grid by:
- Warping: We transform candidate regions into a standardized square grid using perspective warping.
- Sampling: To avoid border noise, we sample only the center area of each color cell.
- Averaging: We calculate the observed color as the mean RGB value of these sampled pixels.
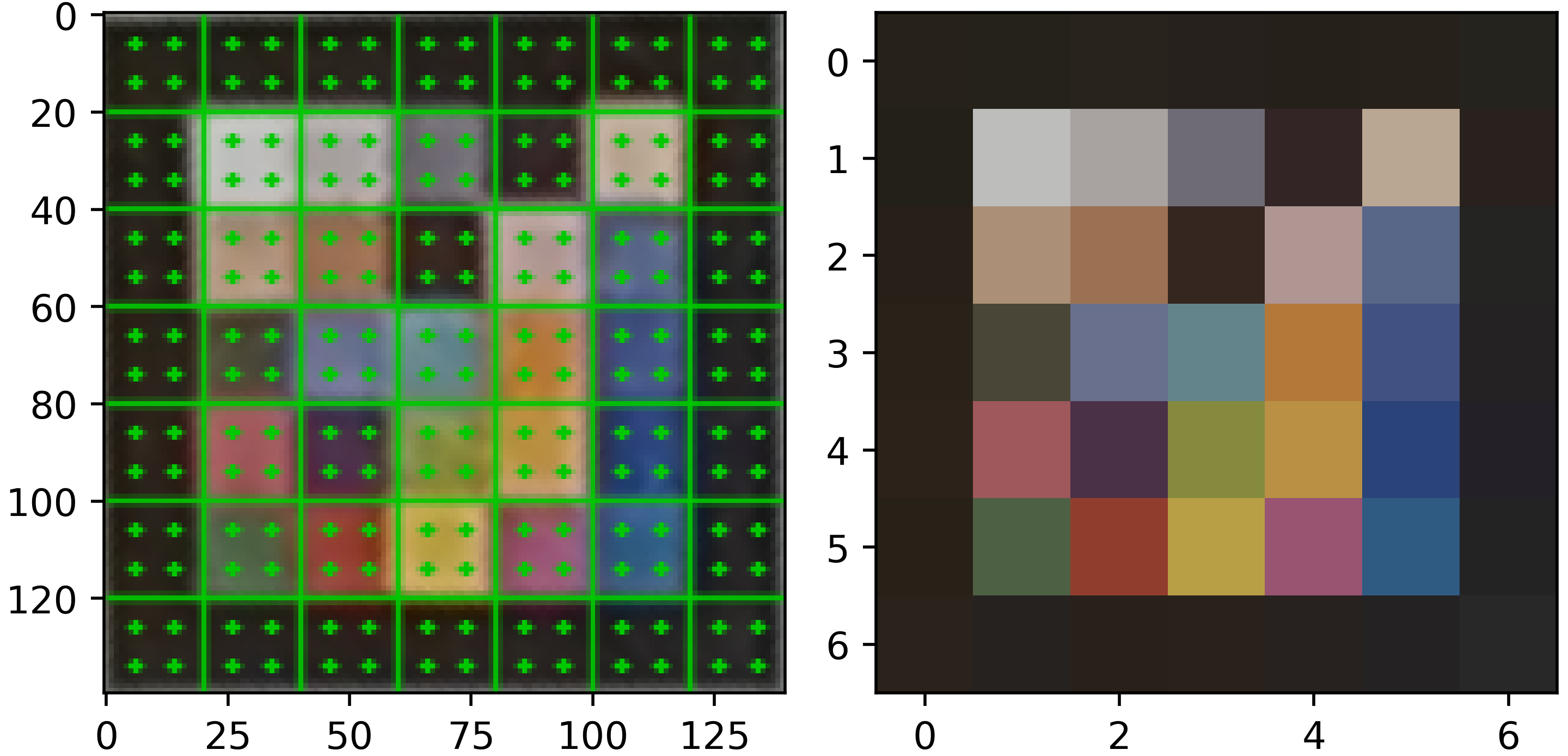 |
|---|
Left: result of the marker wraping. Center: result of the central cell color averaging. Right: reference colors.
Color Transformation Learning:
We fit a linear regression model for each image to find the function that minimizes the distance between observed and reference colors :
Where is the transformation matrix and is the offset vector.
Color Space and Method Selection
To optimize the color correction framework, we evaluated several design variations. For marker design, we compared 5×5 and 6×6 grids and with different colors. We finally selected the 5×5 configuration (25 patches) as it provided sufficient color gamut coverage while remaining practical for clinical use. Regarding color spaces, we tested transformations in RGB, Linear RGB, and CIELAB, ultimately selecting standard RGB for its favorable balance between performance and processing latency. For the transformation method, we chose linear regression over more complex polynomial mappings to prevent color artifacting and ensure consistent results across different camera sensors.
Performance Results
The error metric used is the standard color difference metric (Delta E) (CIEDE2000) in the CIELAB color space to account for a perceptually meaningful measure of color accuracy. The success criterion is set as , which ensures the color difference is not significant to the human eye.
| Metric | Result | Success Criterion | Outcome |
|---|---|---|---|
| 4.35 (4.29-4.42) | ≤ 5.0 | PASS |
Verification and Validation Protocol
Test Design:
- Images captured with identical markers under "warm", "neutral", and "cool" lighting conditions.
- Evaluation of color correction performance using the perceptual color difference (Delta E) metric.
- Visual assessment of corrected images to ensure clinical interpretability and absence of artifacts.
Complete Test Protocol:
- Input: RGB images with reference markers.
- Processing: Automatic detection, perspective rectification, and linear mapping calculation.
- Output: Color-corrected image and mapping error metrics.
- Reference standard: Theoretical patch values.
- Statistical analysis: Mean calculated with 95% confidence intervals via bootstrapping (5000 iterations).
Data Analysis Methods:
- calculation using the CIEDE2000 formula to quantify the perceptual difference between the corrected result and reality.
Test Conclusions:
- The model successfully met the predefined success criterion (), indicating that the residual color error is not significant to the human eye. This ensures high fidelity for downstream clinical sign quantification.
Bias Analysis and Fairness Evaluation
Objective: Ensure the color correction performs consistently across different lighting environments and marker configurations.
Subpopulation Analysis Protocol:
1. Lighting Temperature Analysis:
- Performance evaluated under diverse illumination conditions: warm, neutral, and cool lighting.
- Success criterion: (not significant to human eye).
| Lighting Temperature | Num. Images | Mean | Outcome |
|---|---|---|---|
| Warm | 5 | 4.39 (4.31-4.50) | PASS |
| Neutral | 5 | 4.31 (4.21-4.40) | PASS |
| Cool | 5 | 4.36 (4.24-4.48) | PASS |
Results Summary:
- The method demonstrated reliable color correction performance across different lighting temperatures, meeting all success criteria.
- The method's performance is robust to variations in illumination conditions, indicating its generalizability across diverse clinical photography scenarios.
2. Marker Configuration Analysis:
- Performance when using different number of markers (from 1 to 6).
- Success criterion: (not significant to human eye).
| Num. Markers | Num. Images | Mean | Outcome |
|---|---|---|---|
| 1 | 15 | 4.61 (4.54-4.69) | PASS |
| 2 | 15 | 4.45 (4.42-4.48) | PASS |
| 3 | 15 | 4.39 (4.37-4.41) | PASS |
| 4 | 15 | 4.37 (4.34-4.39) | PASS |
| 5 | 15 | 4.35 (4.32-4.38) | PASS |
| 6 | 15 | 4.35 (4.29-4.42) | PASS |
Results Summary:
- The method demonstrated reliable color correction performance across different marker configurations, meeting all success criteria.
- Although the performance is robust even with a single marker, the use of multiple markers provides a more accurate and robust color transformation.
- The use of multiple markers allows the method to account for spatial lighting variations, which can further enhance the accuracy of the color correction.
Bias Mitigation Strategies:
- Marker filtering: We filter out marker candidates that do not present the expected quality criteria (e.g., presence of a black border, color content, light reflections) to ensure that only valid markers are used for color correction, which helps mitigate errors due to misidentification.
- Multiple color patches: The use of a 5x5 grid of color patches provides a broad color gamut coverage, allowing for a more accurate and robust color transformation that can account for variations in lighting and camera sensors.
- Linear transformation: By using a linear regression model to learn the color transformation, we can effectively correct for systematic color shifts while avoiding overfitting that could occur with more complex models, which helps ensure consistent performance across different images and conditions.
- Multi-Marker Support: The algorithm can aggregate color data from multiple markers to account for spatial lighting variations across the body.
Bias Analysis Conclusion:
- The color correction framework demonstrates robust performance across different lighting conditions and marker configurations.
- The method's performance is within acceptable limits across all subpopulations.
Body Site Identification
Model Overview
Reference: R-TF-028-001 AI/ML Description - Body Site Identification section
This model identifies anatomical locations from images to support site-specific scoring systems and bias monitoring.
Clinical Significance: Anatomical site identification enables automatic application of site-specific scoring rules and demographic performance monitoring.
Data Requirements and Annotation
Data Requirements: Images annotated with anatomical location
Model-specific annotation: Body Site Identification (R-TF-028-024 Data Annotation Instructions)
Medical experts label images with:
- Fine-grained anatomical sites (e.g., dorsal hand, volar forearm)
- Body regions (head/neck, trunk, upper extremity, lower extremity)
- Consensus for ambiguous cases
Dataset statistics:
- Images with body site annotations: 61023 images
- Training set: 80% of the images plus 10% of healthy skin images
- Validation set: 10% of the images
- Test set: 20% of the images
Training Methodology
Architecture: EfficientNet-B2, a convolutional neural network optimized for image classification tasks with a final layer adapted for a binary output.
- Transfer learning from pre-trained weights (ImageNet)
- Input size: RGB images at 272 pixels resolution
Other architectures and resolutions were evaluated during model selection, with EfficientNet-B2 at 272x272 pixels providing the best balance of performance and computational efficiency. Lower resolutions led to loss of detail, while higher resolutions increased computational cost without significant performance gains. Vision Transformer architectures were also evaluated, showing lower performance likely due to the limited dataset size for this specific task.
Training approach:
- Pre-processing: Normalization of input images to standard mean and std of the ImageNet dataset. Other normalizations were evaluated during model selection, with ImageNet normalization providing the best performance.
- Data augmentation: Rotations, mirroring, color jittering, cropping, zoom-out, brightness/contrast adjustments, blur. The global color changes introduced by some augmentations (e.g., color jittering, brightness/contrast adjustments) were carefully tuned to avoid altering the visual appearance. A global augmentation intensity was evaluated to reduce overfitting while preserving the clinical sign characteristics and model performance.
- Data sampler: Batch size 64, with balanced sampling to ensure uniform class distribution across intensity levels. Larger and smaller batch sizes were evaluated during model selection, with non-significant performance differences observed.
- Backbone architecture: An DeepLabV3+ segmentation head was added on top of the EfficientNet-B2 backbone to perform pixel-wise segmentation. Other segmentation heads were evaluated during model selection (e.g., U-Net, FCN), with DeepLabV3+ providing the best performance likely due to its atrous spatial pyramid pooling module that captures multi-scale context.
- Loss function: Combined Cross-entropy loss with logits and Jaccard loss. Associated weights were set based on a hyperparameter search. Weighted cross-entropy loss was evaluated during model selection, with no significant performance differences observed, as the balanced sampling strategy provided sufficient class balance to avoid the need for weighted loss.
- Optimizer: AdamW with learning rate 0.001, betas (0.9, 0.999), weight decay 0. SGD and RMSProp optimizers were evaluated during model selection, with Adam providing the best convergence speed and final performance, likely due to the dataset size and complexity.
- Training duration: 400 epochs. At this point, the model had fully converged with evaluating metrics on the validation set stabilizing.
- Learning rate scheduler: StepLR with step size 1 epoch, and gamma to decay the learning rate to 1.e-2 the starting learning rate at the end of training. Other schedulers were evaluated during model selection (e.g., cosine annealing, ReduceLROnPlateau), with no significant performance differences observed.
- Evaluation metrics: accuracy, F1-score, sensitivity, and specificity calculated on the validation set after each epoch to monitor training progress and select the best model based on validation IoU.
- Model freezing: No freezing of layers was applied. Freezing strategies were evaluated during model selection, showing a negative impact on performance likely due to the domain gap between ImageNet and dermatology images.
Post-processing:
- Sigmoid activation to obtain probability distributions
- Binary classification thresholds to convert probabilities to binary masks.
Performance Results
Performance evaluated using Balanced Accuracy (BAcc) to account for class imbalance across anatomical sites.
Success criteria:
| Metric | Result | Success Criterion | Outcome |
|---|---|---|---|
| BAcc. Buttock | 0.737 (0.678, 0.786) | ≥ 0.70 | PASS |
| BAcc. Arm | 0.855 (0.842, 0.869) | ≥ 0.70 | PASS |
| BAcc. Armpit | 0.831 (0.795, 0.868) | ≥ 0.70 | PASS |
| BAcc. Hand | 0.973 (0.969, 0.977) | ≥ 0.70 | PASS |
| BAcc. Hand nail | 0.916 (0.901, 0.929) | ≥ 0.70 | PASS |
| BAcc. Genitals | 0.921 (0.904, 0.942) | ≥ 0.70 | PASS |
| BAcc. Leg | 0.869 (0.854, 0.883) | ≥ 0.70 | PASS |
| BAcc. Knee | 0.77 (0.738, 0.804) | ≥ 0.70 | PASS |
| BAcc. Foot | 0.907 (0.893, 0.923) | ≥ 0.70 | PASS |
| BAcc. Foot nail | 0.906 (0.887, 0.934) | ≥ 0.70 | PASS |
| BAcc. Toe | 0.935 (0.922, 0.952) | ≥ 0.70 | PASS |
| BAcc. Finger | 0.942 (0.932, 0.952) | ≥ 0.70 | PASS |
| BAcc. Close-up image | 0.901 (0.894, 0.91) | ≥ 0.70 | PASS |
| BAcc. Top of the head | 0.818 (0.784, 0.86) | ≥ 0.70 | PASS |
| BAcc. Scalp | 0.882 (0.86, 0.906) | ≥ 0.70 | PASS |
| BAcc. Back of the head | 0.754 (0.715, 0.799) | ≥ 0.70 | PASS |
| BAcc. Face | 0.939 (0.933, 0.944) | ≥ 0.70 | PASS |
| BAcc. Mouth | 0.959 (0.952, 0.966) | ≥ 0.70 | PASS |
| BAcc. Tongue | 0.851 (0.801, 0.898) | ≥ 0.70 | PASS |
| BAcc. Ear | 0.91 (0.895, 0.925) | ≥ 0.70 | PASS |
| BAcc. Eye | 0.908 (0.894, 0.918) | ≥ 0.70 | PASS |
| BAcc. Nose | 0.959 (0.953, 0.965) | ≥ 0.70 | PASS |
| BAcc. Neck | 0.839 (0.822, 0.858) | ≥ 0.70 | PASS |
| BAcc. Back | 0.848 (0.825, 0.871) | ≥ 0.70 | PASS |
| BAcc. Trunk | 0.859 (0.844, 0.875) | ≥ 0.70 | PASS |
Verification and Validation Protocol
Test Design:
- Test set with expert-confirmed anatomical locations
- Coverage of all major body sites
- Challenging cases: Similar sites, unusual angles, partial views
- Evaluation across diverse Fitzpatrick skin types and severity levels
Complete Test Protocol:
- Input: RGB images from test set with expert erythema intensity annotations
- Processing: Model inference with probability distribution output
- Output: Predicted body sites with confidence scores
- Statistical analysis: BAcc Accuracy, Recall and Precision with Confidence Intervals calculated using bootstrap resampling (2000 iterations).
- Robustness checks was performed to ensure consistent performance across several image transformations that do not alter the clinical sign appearance and simulate real-world variations (rotations, brightness/contrast adjustments, zoom, and image quality).
Data Analysis Methods:
- BAcc calculation with Confidence Intervals
- Bootstrap resampling (2000 iterations) for 95% confidence intervals
Test Conclusions:
The model's classification performance, rigorously assessed using the BAcc metric across all anatomical sites, successfully and robustly meets the predefined Success Criterion of . A review of the results reveals that every single anatomical site demonstrates a lower bound of the 95% CI that is above the Success Criterion of 0.70, thus satisfying the PASS criterion with a high degree of confidence and indicating minimal variability below the acceptable threshold. For instance, the site with the lowest performance, "Buttock," still registered a lower CI of , and the mean BAcc (0.737) and the upper bound of the CI (0.786) are both well above the criterion. Conversely, regions such as "Hand" () and "Nose" () demonstrate exceptional performance, with their lower CI bounds still exceeding . This comprehensive and uniform success confirms that the model maintains a high and reliable degree of classification accuracy across a diverse range of body locations, validating its robust performance and indicating no discernible bias with respect to anatomical site.
Bias Analysis and Fairness Evaluation
Objective: Ensure anatomical site identification performs consistently across demographics.
Subpopulation Analysis Protocol:
1. Fitzpatrick Skin Type Analysis (Critical for induration):
- RMAE calculation per Fitzpatrick type (I-II, III-IV, V-VI)
- Comparison of model performance vs. expert inter-observer variability per skin type
- Success criterion: Consistent RMAE across severity levels
Bias Mitigation Strategies:
- Training data balanced across Fitzpatrick types
- Diverse representation of body types
Results Summary:
- Fitzpatrick I-II
| Metric | Result: Mean (95% CI) | # samples | Success Criterion | Outcome |
|---|---|---|---|---|
| BAcc. Buttock | 0.764 (0.689, 0.853) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Arm | 0.827 (0.799, 0.848) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Armpit | 0.832 (0.767, 0.887) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Hand | 0.948 (0.935, 0.960) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Hand nail | 0.903 (0.879, 0.927) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Genitals | 0.899 (0.869, 0.930) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Leg | 0.848 (0.821, 0.875) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Knee | 0.738 (0.674, 0.807) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Foot | 0.893 (0.867, 0.918) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Foot nail | 0.875 (0.841, 0.923) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Toe | 0.917 (0.891, 0.946) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Finger | 0.920 (0.904, 0.937) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Close-up image | 0.920 (0.912, 0.929) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Top of the head | 0.817 (0.772, 0.866) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Scalp | 0.885 (0.856, 0.914) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Back of the head | 0.767 (0.684, 0.852) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Face | 0.948 (0.941, 0.956) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Mouth | 0.972 (0.965, 0.980) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Tongue | 0.897 (0.846, 0.963) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Ear | 0.908 (0.884, 0.935) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Eye | 0.895 (0.875, 0.912) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Nose | 0.963 (0.955, 0.970) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Neck | 0.851 (0.830, 0.872) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Back | 0.805 (0.764, 0.844) | 5100 | BAcc ≥ 0.70 | PASS |
| BAcc. Trunk | 0.858 (0.832, 0.884) | 5100 | BAcc ≥ 0.70 | PASS |
- Fitzpatrick III-IV
| Metric | Result: Mean (95% CI) | # samples | Success Criterion | Outcome |
|---|---|---|---|---|
| BAcc. Buttock | 0.719 (0.635, 0.789) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Arm | 0.866 (0.845, 0.889) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Armpit | 0.863 (0.802, 0.93) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Hand | 0.982 (0.979, 0.986) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Hand nail | 0.918 (0.896, 0.936) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Genitals | 0.941 (0.918, 0.967) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Leg | 0.869 (0.845, 0.891) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Knee | 0.78 (0.725, 0.84) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Foot | 0.893 (0.865, 0.917) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Foot nail | 0.916 (0.88, 0.945) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Toe | 0.935 (0.91, 0.957) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Finger | 0.954 (0.942, 0.968) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Close-up image | 0.871 (0.857, 0.886) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Top of the head | 0.806 (0.72, 0.899) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Scalp | 0.855 (0.814, 0.899) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Back of the head | 0.72 (0.657, 0.793) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Face | 0.927 (0.915, 0.936) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Mouth | 0.944 (0.927, 0.958) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Tongue | 0.806 (0.737, 0.884) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Ear | 0.91 (0.886, 0.933) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Eye | 0.922 (0.903, 0.938) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Nose | 0.952 (0.94, 0.966) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Neck | 0.824 (0.797, 0.856) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Back | 0.858 (0.826, 0.889) | 5633 | BAcc ≥ 0.70 | PASS |
| BAcc. Trunk | 0.86 (0.838, 0.883) | 5633 | BAcc ≥ 0.70 | PASS |
- Fitzpatrick VI-VI
| Metric | Result: Mean (95% CI) | # samples | Success Criterion | Outcome |
|---|---|---|---|---|
| BAcc. Buttock | 0.721 (0.555, 0.915) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Arm | 0.862 (0.832, 0.891) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Armpit | 0.735 (0.6, 0.874) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Hand | 0.955 (0.942, 0.971) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Hand nail | 0.93 (0.902, 0.961) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Genitals | 0.927 (0.876, 0.968) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Leg | 0.877 (0.854, 0.904) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Knee | 0.773 (0.712, 0.832) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Foot | 0.946 (0.92, 0.968) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Foot nail | 0.933 (0.896, 0.97) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Toe | 0.965 (0.939, 0.987) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Finger | 0.95 (0.93, 0.969) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Close-up image | 0.837 (0.8, 0.873) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Top of the head | 0.84 (0.735, 0.945) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Scalp | 0.92 (0.877, 0.957) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Back of the head | 0.785 (0.718, 0.866) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Face | 0.918 (0.895, 0.941) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Mouth | 0.913 (0.881, 0.952) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Tongue | 0.849 (0.713, 0.999) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Ear | 0.912 (0.877, 0.948) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Eye | 0.903 (0.868, 0.94) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Nose | 0.951 (0.923, 0.978) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Neck | 0.834 (0.779, 0.878) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Back | 0.893 (0.853, 0.932) | 1476 | BAcc ≥ 0.70 | PASS |
| BAcc. Trunk | 0.856 (0.82, 0.898) | 1476 | BAcc ≥ 0.70 | PASS |
Bias Analysis Conclusion:
The model's classification performance, assessed by BAcc across various anatomical sites and all Fitzpatrick scale categories, successfully meets the predefined Success Criterion of . Focusing on the most stringent compliance, every single site across all three Fitzpatrick groups demonstrates a mean BAcc that is above the Success Criterion of 0.70, thus robustly satisfying the PASS criterion. For example, in the Fitzpatrick I-II group, the lowest mean BAcc is for "Buttock" at 0.764 (95% CI: 0.689, 0.853), where the upper bound of the CI (0.853) and the mean BAcc (0.764) are both well above the criterion. Regions like "Hand" and "Face" consistently exhibit the highest performance across all groups, frequently registering mean BAcc values above (e.g., for Face in Fitzpatrick I-II) and with their entire 95% CI well above the 0.70 threshold. This comprehensive performance demonstrates that the model maintains a high degree of classification accuracy across different anatomical locations and a broad spectrum of skin tones, indicating minimal bias with respect to anatomical site and Fitzpatrick scale category.
Summary and Conclusion
The development and validation activities described in this report provide objective evidence that the AI algorithms for Legit.Health Plus meet their predefined specifications and performance requirements.
Status of model development and validation:
- ICD Category Distribution and Binary Indicators: [Status to be updated]
- Visual Sign Intensity Models: [Status to be updated]
- Lesion Quantification Models: [Status to be updated]
- Surface Area Models: [Status to be updated]
- Non-Clinical Support Models: [Status to be updated]
The development process adhered to the company's QMS and followed Good Machine Learning Practices. Models meeting their success criteria are considered verified, validated, and suitable for release and integration into the Legit.Health Plus medical device.
State of the Art Compliance and Development Lifecycle
Software Development Lifecycle Compliance
The AI models in Legit.Health Plus were developed in accordance with state-of-the-art software development practices and international standards:
Applicable Standards and Guidelines:
- IEC 62304:2006+AMD1:2015 - Medical device software lifecycle processes
- ISO 13485:2016 - Quality management systems for medical devices
- ISO 14971:2019 - Application of risk management to medical devices
- ISO/IEC 25010:2011 - Systems and software quality requirements and evaluation (SQuaRE)
- FDA Guidance on Software as a Medical Device (SAMD) - Clinical evaluation and predetermined change control plans
- IMDRF/SaMD WG/N41 FINAL:2017 - Software as a Medical Device: Key Definitions
- Good Machine Learning Practice (GMLP) - FDA/Health Canada/UK MHRA Guiding Principles (2021)
- Proposed Regulatory Framework for Modifications to AI/ML-Based SaMD - FDA Discussion Paper (2019)
Development Lifecycle Phases Implemented:
- Requirements Analysis: Comprehensive AI model specifications defined in
R-TF-028-001 AI/ML Description - Development Planning: Structured development plan in
R-TF-028-002 AI Development Plan - Risk Management: AI-specific risk analysis in
R-TF-028-011 AI Risk Matrix - Design and Architecture: State-of-the-art architectures (Vision Transformers, CNNs, object detection, segmentation)
- Implementation: Following coding standards and version control practices
- Verification: Unit testing, integration testing, and algorithm validation
- Validation: Clinical performance testing against predefined success criteria
- Release: Version-controlled releases with complete traceability
- Maintenance: Post-market surveillance and performance monitoring
Version Control and Traceability:
- All model versions tracked in version control systems (Git)
- Complete traceability from requirements to validation results
- Dataset versions documented with checksums and provenance
- Model artifacts stored with complete training metadata
- Documented change control process for model updates
State of the Art in AI Development
Best Practices Implemented:
1. Data Management Excellence:
- Multi-source data collection with demographic diversity
- Rigorous data quality control and curation processes
- Systematic annotation protocols with multi-expert consensus
- Data partitioning strategies preventing data leakage
- Sequestered test sets for unbiased evaluation
2. Model Architecture Selection:
- Use of state-of-the-art architectures (Vision Transformers for classification, YOLO/Faster R-CNN for detection, U-Net/DeepLab for segmentation)
- Transfer learning from large-scale pre-trained models
- Architecture selection based on published benchmark performance
- Justification of architecture choices documented per model
3. Training Best Practices:
- Systematic hyperparameter optimization
- Cross-validation and early stopping to prevent overfitting
- Data augmentation for robustness and generalization
- Multi-task learning where clinically appropriate
- Monitoring of training metrics and convergence
4. Model Calibration and Post-Processing:
- Temperature scaling for probability calibration
- Test-time augmentation for robust predictions
- Ensemble methods where applicable
- Uncertainty quantification for model predictions
5. Comprehensive Validation:
- Independent test sets never used during development
- External validation on diverse datasets
- Clinical reference standard from expert consensus
- Statistical rigor with confidence intervals
- Comprehensive subpopulation analysis
6. Bias Mitigation and Fairness:
- Systematic bias analysis across demographic subpopulations
- Fitzpatrick skin type stratification in all analyses
- Data collection strategies ensuring demographic diversity
- Bias monitoring models (DIQA, Fitzpatrick identification)
- Transparent reporting of performance disparities
7. Explainability and Transparency:
- Attention visualization for model interpretability (where applicable)
- Clinical reasoning transparency (top-k predictions with probabilities)
- Documentation of model limitations and known failure modes
- Clear communication of uncertainty in predictions
Risk Management Throughout Lifecycle
Risk Management Process:
Risk management is integrated throughout the entire AI development lifecycle following ISO 14971:
1. Risk Analysis:
- Identification of AI-specific hazards (data bias, model errors, distribution shift)
- Hazardous situation analysis (incorrect predictions leading to clinical harm)
- Risk estimation combining probability and severity
2. Risk Evaluation:
- Comparison of risks against predefined acceptability criteria
- Benefit-risk analysis for each AI model
- Clinical impact assessment of potential errors
3. Risk Control:
- Inherent safety by design (offline models, no learning from deployment data)
- Protective measures (DIQA filtering, domain validation, confidence thresholds)
- Information for safety (user training, clinical decision support context)
4. Residual Risk Evaluation:
- Assessment of risks after control measures
- Verification that all risks reduced to acceptable levels
- Overall residual risk acceptability
5. Risk Management Review:
- Production and post-production information review
- Update of risk management file
- Traceability to safety risk matrix (
R-TF-028-011 AI Risk Matrix)
AI-Specific Risk Controls:
- Data Quality Risks: Multi-source collection, systematic annotation, quality control
- Model Overfitting: Sequestered test sets, cross-validation, regularization
- Bias and Fairness: Demographic diversity, subpopulation analysis, bias monitoring
- Model Uncertainty: Calibration, confidence scores, uncertainty quantification
- Distribution Shift: Domain validation, DIQA filtering, performance monitoring
- Clinical Misinterpretation: Clear communication, clinical context, user training
Information Security
Cybersecurity Considerations:
The AI models are designed with information security principles integrated throughout development:
1. Model Security:
- Model parameters stored securely with access controls
- Model integrity verification (checksums, digital signatures)
- Protection against model extraction or reverse engineering
- Secure deployment pipelines
2. Data Security:
- Patient data protection throughout development (de-identification, anonymization)
- Secure data storage with encryption at rest
- Secure data transmission with encryption in transit
- Access controls and audit logging for training data
3. Inference Security:
- Secure API endpoints for model inference
- Input validation to prevent adversarial attacks
- Rate limiting and authentication
- Output validation and sanity checking
4. Privacy Considerations:
- No patient-identifiable information stored in models
- Training data anonymization and de-identification
- Compliance with GDPR, HIPAA, and applicable privacy regulations
- Data minimization principles applied
5. Vulnerability Management:
- Regular security assessments of AI infrastructure
- Dependency scanning for software libraries
- Patch management for underlying frameworks
- Incident response procedures
6. Adversarial Robustness:
- Consideration of adversarial attack scenarios
- Input preprocessing to detect anomalous inputs
- Domain validation to reject out-of-distribution inputs
- DIQA filtering to reject manipulated or low-quality images
Cybersecurity Risk Assessment:
Cybersecurity risks are addressed in the overall device risk management file, including:
- Threat modeling for AI components
- Attack surface analysis
- Mitigation strategies and security controls
- Monitoring and incident response
Verification and Validation Strategy
Verification Activities (confirming that the AI models implement their specifications):
- Code reviews and static analysis
- Unit testing of model components
- Integration testing of model pipelines
- Architecture validation against specifications
- Performance benchmarking against target metrics
Validation Activities (confirming that AI models meet intended use):
- Independent test set evaluation with sequestered data
- External validation on diverse datasets
- Clinical reference standard comparison
- Subpopulation performance analysis
- Real-world performance assessment
- Usability and clinical workflow validation
Documentation of Verification and Validation:
Complete documentation is maintained for all verification and validation activities:
- Test protocols with detailed methodology
- Complete test results with statistical analysis
- Data summaries and test conclusions
- Traceability from requirements to test results
- Identified deviations and their resolutions
This comprehensive approach ensures compliance with GSPR 17.2 requirements for software development in accordance with state of the art, incorporating development lifecycle management, risk management, information security, verification, and validation.
Integration Verification Package
To ensure that the AI models produce identical outputs when integrated into the Legit.Health Plus software environment as they did during development and validation, an Integration Verification Package has been prepared for each model in accordance with GP-028 AI Development.
Purpose
The Integration Verification Package enables the Software Development team to:
- Verify that models are correctly integrated without alterations to their inference behavior
- Detect any environment discrepancies that could affect model outputs
- Provide objective evidence of output equivalence between development and production environments
- Support regulatory compliance by demonstrating traceability between development validation and deployed system verification per IEC 62304
Package Location and Structure
All Integration Verification Packages are stored in the secure, version-controlled S3 bucket with the following structure:
s3://legit-health-plus/integration-verification/
├── icd-category-distribution/
│ ├── images/
│ ├── expected_outputs.csv
│ └── manifest.json
├── erythema-intensity/
│ ├── images/
│ ├── expected_outputs.csv
│ └── manifest.json
├── desquamation-intensity/
│ ├── images/
│ ├── expected_outputs.csv
│ └── manifest.json
├── induration-intensity/
│ ├── images/
│ ├── expected_outputs.csv
│ └── manifest.json
├── pustule-intensity/
│ ├── images/
│ ├── expected_outputs.csv
│ └── manifest.json
├── [... additional models ...]
├── diqa/
│ ├── images/
│ ├── expected_outputs.csv
│ └── manifest.json
└── domain-validation/
├── images/
├── expected_outputs.csv
└── manifest.json
Package Contents Per Model
For each AI model in the Legit.Health Plus device, the Integration Verification Package includes:
Reference Test Images
- Location:
s3://legit-health-plus/integration-verification/{MODEL_NAME}/images/ - Content: A curated subset of images from the model's held-out test set
- Selection Criteria: Images representative of the model's input domain, including diverse conditions, demographics, and imaging modalities
- Format: Original image format (JPEG/PNG) without additional processing
Expected Outputs File
- Location:
s3://legit-health-plus/integration-verification/{MODEL_NAME}/expected_outputs.csv - Schema:
| Column | Type | Description |
|---|---|---|
image_id | string | Unique identifier matching the image filename |
expected_output | string/float | Model's expected output (JSON-encoded for complex outputs) |
output_type | string | Output category: classification_probability, regression_value, segmentation_mask_hash, detection_boxes |
preprocessing_hash | string | SHA-256 hash of the preprocessed input tensor |
- Generation: Outputs generated from the validated development model using the exact configuration documented in this report
Verification Manifest
- Location:
s3://legit-health-plus/integration-verification/{MODEL_NAME}/manifest.json - Contents:
{
"model_name": "erythema-intensity",
"model_version": "1.0.0",
"package_version": "1.0.0",
"creation_timestamp": "2026-01-27T10:00:00Z",
"created_by": "AI Team",
"num_test_images": 100,
"model_weights_sha256": "abc123...",
"preprocessing": {
"resize": [272, 272],
"normalization": "imagenet",
"color_space": "RGB"
},
"acceptance_criteria": {
"metric": "output_tolerance",
"tolerance": 1e-5,
"pass_rate_required": 1.0
},
"development_report_reference": "R-TF-028-005 v1.0"
}
Acceptance Criteria
The following acceptance criteria apply to integration verification:
| Model Type | Metric | Acceptance Criterion |
|---|---|---|
| Classification (ICD, Binary Indicators) | Probability difference | ε ≤ 1e-5 per class |
| Intensity Quantification | Output score difference | ε ≤ 1e-5 |
| Segmentation | Mask IoU | ≥ 0.9999 |
| Detection | Box IoU + class match | IoU ≥ 0.9999, exact class match |
| Quality Assessment (DIQA) | Score difference | ε ≤ 1e-5 |
Overall Pass Criterion: 100% of test images must meet the acceptance criteria for the integration verification to pass.
Model-Specific Package Details
The following table summarizes the Integration Verification Package for each model:
Clinical Models - ICD Classification and Binary Indicators
| Model | Output Type | Storage Path |
|---|---|---|
| ICD Category Distribution | Classification probabilities (346 classes) | icd-category-distribution/ |
| Binary Indicators | 6 probability scores | icd-category-distribution/ |
Clinical Models - Visual Sign Intensity Quantification
| Model | Output Type | Storage Path |
|---|---|---|
| Erythema Intensity | Regression (0-9 scale) | erythema-intensity/ |
| Desquamation Intensity | Regression (0-9 scale) | desquamation-intensity/ |
| Induration Intensity | Regression (0-9 scale) | induration-intensity/ |
| Pustule Intensity | Regression (0-9 scale) | pustule-intensity/ |
| Crusting Intensity | Regression (0-9 scale) | crusting-intensity/ |
| Xerosis Intensity | Regression (0-9 scale) | xerosis-intensity/ |
| Swelling Intensity | Regression (0-9 scale) | swelling-intensity/ |
| Oozing Intensity | Regression (0-9 scale) | oozing-intensity/ |
| Excoriation Intensity | Regression (0-9 scale) | excoriation-intensity/ |
| Lichenification Intensity | Regression (0-9 scale) | lichenification-intensity/ |
Clinical Models - Wound Characteristic Assessment
| Model | Output Type | Storage Path |
|---|---|---|
| Wound Edge: Diffused | Binary classification | wound-edge-diffused/ |
| Wound Edge: Thickened | Binary classification | wound-edge-thickened/ |
| Wound Edge: Delimited | Binary classification | wound-edge-delimited/ |
| Wound Edge: Indistinguishable | Binary classification | wound-edge-indistinguishable/ |
| Wound Edge: Damaged | Binary classification | wound-edge-damaged/ |
| Wound Tissue: Bone | Binary classification | wound-tissue-bone/ |
| Wound Tissue: Subcutaneous | Binary classification | wound-tissue-subcutaneous/ |
| Wound Tissue: Muscle | Binary classification | wound-tissue-muscle/ |
| Wound Tissue: Intact | Binary classification | wound-tissue-intact/ |
| Wound Tissue: Dermis-Epidermis | Binary classification | wound-tissue-dermis-epidermis/ |
| Wound Bed: Necrotic | Binary classification | wound-bed-necrotic/ |
| Wound Bed: Closed | Binary classification | wound-bed-closed/ |
| Wound Bed: Granulation | Binary classification | wound-bed-granulation/ |
| Wound Bed: Epithelial | Binary classification | wound-bed-epithelial/ |
| Wound Bed: Slough | Binary classification | wound-bed-slough/ |
| Wound Exudate: Serous | Binary classification | wound-exudate-serous/ |
| Wound Exudate: Fibrinous | Binary classification | wound-exudate-fibrinous/ |
| Wound Exudate: Purulent | Binary classification | wound-exudate-purulent/ |
| Wound Exudate: Bloody | Binary classification | wound-exudate-bloody/ |
| Perilesional Erythema | Binary classification | perilesional-erythema/ |
| Perilesional Maceration | Binary classification | perilesional-maceration/ |
| Biofilm Tissue | Binary classification | biofilm-tissue/ |
| Wound Stage Classification | Multi-class (6 stages) | wound-stage/ |
| Wound Intensity (AWOSI) | Regression (0-10 scale) | wound-awosi/ |
Clinical Models - Lesion Quantification
| Model | Output Type | Storage Path |
|---|---|---|
| Inflammatory Nodular Lesion | Detection (bounding boxes + count) | inflammatory-nodular/ |
| Acneiform Lesion Types | Multi-class detection (5 classes) | acneiform-lesion-types/ |
| Inflammatory Lesion | Detection (bounding boxes + count) | inflammatory-lesion/ |
| Hive Lesion | Detection (bounding boxes + count) | hive-lesion/ |
| Nail Lesion Surface | Segmentation mask | nail-lesion-surface/ |
Clinical Models - Surface Area Quantification
| Model | Output Type | Storage Path |
|---|---|---|
| Wound Bed Surface | Segmentation mask | wound-bed-surface/ |
| Wound Granulation Surface | Segmentation mask | wound-granulation-surface/ |
| Wound Biofilm/Slough Surface | Segmentation mask | wound-biofilm-surface/ |
| Wound Necrosis Surface | Segmentation mask | wound-necrosis-surface/ |
| Wound Maceration Surface | Segmentation mask | wound-maceration-surface/ |
| Wound Orthopedic Material Surface | Segmentation mask | wound-orthopedic-surface/ |
| Wound Bone/Cartilage/Tendon Surface | Segmentation mask | wound-bone-surface/ |
| Hair Loss Surface | Segmentation mask | hair-loss-surface/ |
| Hypopigmentation/Depigmentation | Segmentation mask | hypopigmentation-surface/ |
| Hyperpigmentation Surface | Segmentation mask | hyperpigmentation-surface/ |
| Erythema Surface | Segmentation mask | erythema-surface/ |
Clinical Models - Pattern Identification
| Model | Output Type | Storage Path |
|---|---|---|
| Acneiform Inflammatory Pattern | Regression (IGA 0-4 scale) | acneiform-pattern/ |
| Follicular and Inflammatory Pattern | Multi-class (Hurley stages) | follicular-inflammatory-pattern/ |
| Inflammatory Pattern | Classification | inflammatory-pattern/ |
| Inflammatory Pattern Indicator | Binary classification | inflammatory-pattern-indicator/ |
Non-Clinical Models
| Model | Output Type | Storage Path |
|---|---|---|
| DIQA | Quality score (0-1) | diqa/ |
| Domain Validation | Classification (3 classes) | domain-validation/ |
| Skin Surface Segmentation | Segmentation mask | skin-surface-segmentation/ |
| Body Surface Segmentation | Segmentation mask | body-surface-segmentation/ |
| Head Detection | Detection (bounding boxes) | head-detection/ |
Verification Procedure for Software Integration Team
The Software Development team shall follow this procedure after model integration:
-
Environment Preparation:
- Configure the integration environment with dependencies specified in
R-TF-028-006 AI Release Report - Download the Integration Verification Package from S3
- Verify package integrity using manifest checksums
- Configure the integration environment with dependencies specified in
-
Inference Execution:
- Process all reference test images through the integrated model
- Record outputs in the same format as
expected_outputs.csv - Document runtime environment configuration
-
Output Comparison:
- Compare actual outputs against expected outputs using acceptance criteria
- Calculate match rate for each image
- Flag any discrepancies
-
Results Documentation:
- Generate Integration Verification Report including:
- Test execution date and environment details
- Pass/fail status per image
- Overall pass rate
- Any deviations with root cause analysis
- Store report as software verification evidence per IEC 62304
- Generate Integration Verification Report including:
Traceability
| Artifact | Version | Reference |
|---|---|---|
| AI Development Report | 1.0 | This document |
| AI Release Report | 1.0 | R-TF-028-006 |
| Integration Verification Package | 1.0 | S3 bucket |
| Model Weights | Per model | See manifest.json |
The Integration Verification Package version is locked to the corresponding model version and AI Development Report. Any model retraining requires generation of a new Integration Verification Package.
AI Risks Assessment Report
AI Risk Assessment
A comprehensive risk assessment was conducted throughout the development lifecycle in accordance with the R-TF-028-002 AI Development Plan. All identified AI-specific risks related to data, model training, and performance were documented and analyzed in the R-TF-028-011 AI Risk Matrix.
AI Risk Treatment
Control measures were implemented to mitigate all identified risks. Key controls included:
- Rigorous data curation and multi-source collection to mitigate bias.
- Systematic model training and validation procedures to prevent overfitting.
- Use of a sequestered test set to ensure unbiased performance evaluation.
- Implementation of model calibration to improve the reliability of outputs.
Residual AI Risk Assessment
After the implementation of control measures, a residual risk analysis was performed. All identified AI risks were successfully reduced to an acceptable level.
AI Risk and Traceability with Safety Risk
Safety risks related to the AI algorithms (e.g., incorrect assessment suggestion, misinterpretation of data) were identified and traced back to their root causes in the AI development process. These safety risks have been escalated for management in the overall device Safety Risk Matrix, in line with ISO 14971.
Conclusion
The AI development process has successfully managed and mitigated inherent risks to an acceptable level. The benefits of using the Legit.Health Plus algorithms as a clinical decision support tool are judged to outweigh the residual risks.
Related Documents
Project Design and Plan
R-TF-028-001 AI/ML Description- Complete specifications for all AI modelsR-TF-028-002 AI Development Plan- Development methodology and lifecycleR-TF-028-011 AI Risk Matrix- AI-specific risk assessment and mitigation
Data Collection and Annotation
R-TF-028-003 Data Collection Instructions- Public datasets and clinical study data collection protocolsR-TF-028-004 Data Annotation Instructions - ICD-11 Mapping- Foundational clinical label standardization (completed)R-TF-028-004 Data Annotation Instructions - Visual Signs- Intensity, count, and extent annotations for visual sign models (completed)R-TF-028-004 Data Annotation Instructions - DIQA- Image quality assessment annotations (to be created)R-TF-028-004 Data Annotation Instructions - Fitzpatrick- Skin type annotations (to be created)R-TF-028-004 Data Annotation Instructions - Body Site- Anatomical location annotations (if needed)
Signature meaning
The signatures for the approval process of this document can be found in the verified commits at the repository for the QMS. As a reference, the team members who are expected to participate in this document and their roles in the approval process, as defined in Annex I Responsibility Matrix of the GP-001, are:
- Author: Team members involved
- Reviewer: JD-003 Design & Development Manager, JD-004 Quality Manager & Person Responsible for Regulatory Compliance (PRRC)
- Approver: JD-001 General Manager